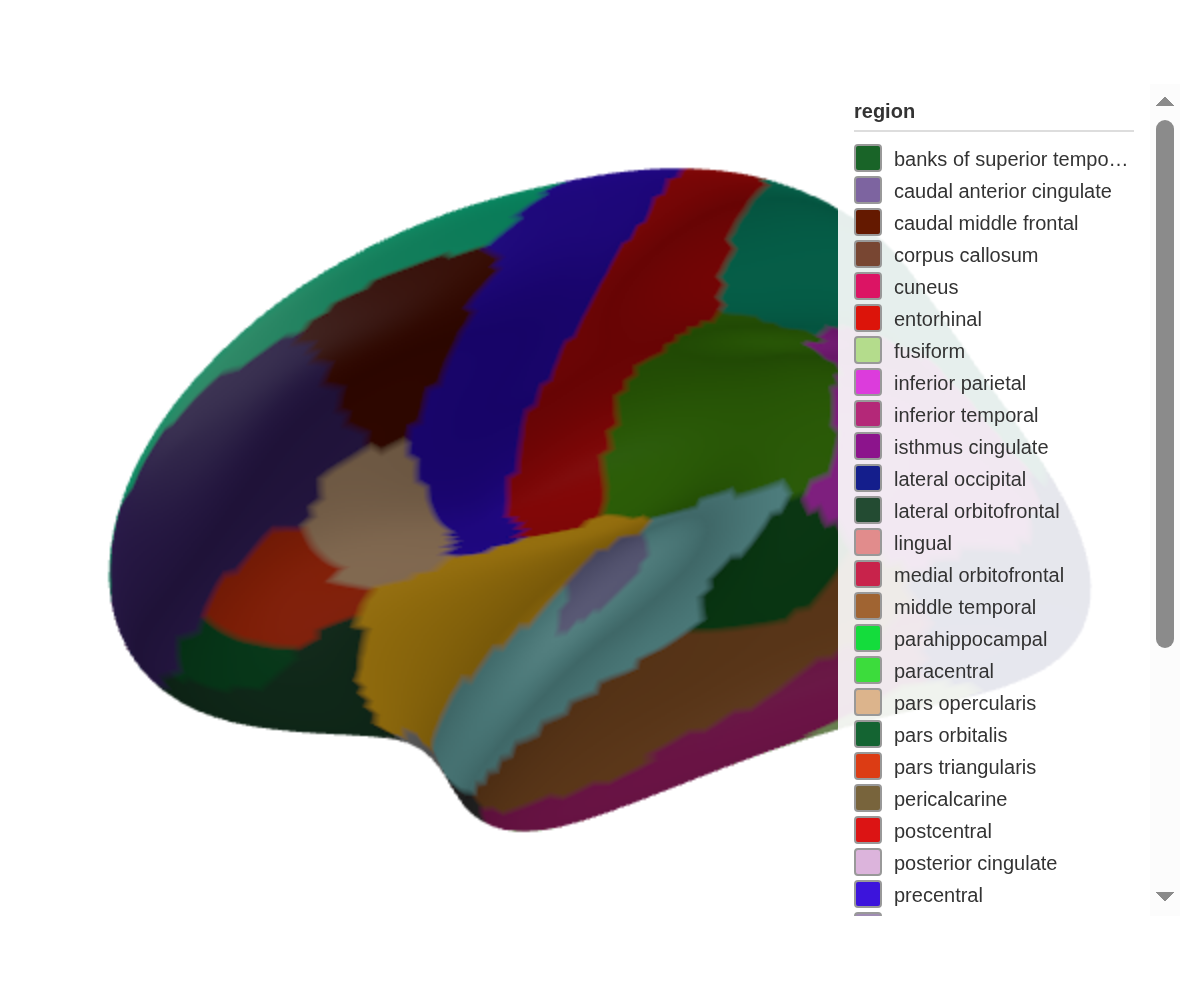
ggseg3d turns brain segmentation data into 3D meshes you can rotate, colour, and export. Version 2.0 replaced the Plotly backend with Three.js — smaller widgets, faster rendering, and a pipe-friendly API that shares atlas objects with the 2D side (ggseg).
This walkthrough covers the full surface area: cortical atlases, subcortical structures, white matter tracts, camera control, colour palettes, edge rendering, and static exports.
Basic usage
One call gets you an interactive brain:
ggseg3d(hemisphere = "left") |>
pan_camera("left lateral")Click and drag to rotate, scroll to zoom, right-click to pan. Hover over any region to see its name.
Camera positions
pan_camera() snaps the camera to standard anatomical
views:
ggseg3d() |>
pan_camera("left lateral")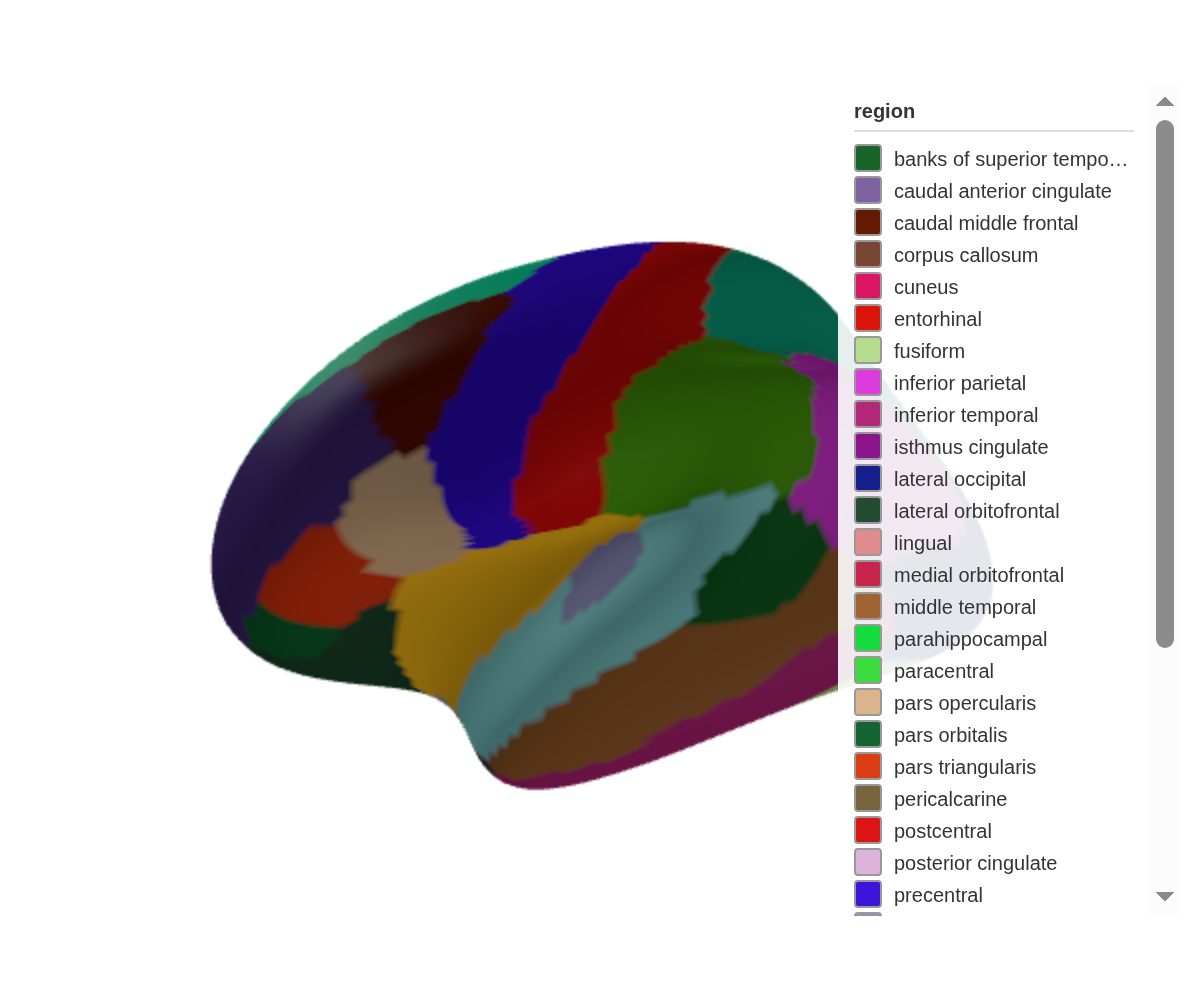
The full set of presets:
-
left lateral,right lateral -
left medial,right medial -
left superior,right superior -
left inferior,right inferior -
left anterior,right anterior -
left posterior,right posterior
For anything else, pass explicit coordinates:
ggseg3d() |>
pan_camera(list(eye = list(x = -200, y = 100, z = 100)))Plotting your data
The core workflow: bring a data frame with a region (or
label) column that matches the atlas, then map a variable
to colour_by. Add text_by to surface values in
the hover tooltip:
some_data <- tibble(
region = c("precentral", "postcentral", "insula", "superior parietal"),
p = c(0.01, 0.04, 0.2, 0.5)
)
ggseg3d(.data = some_data, atlas = dk(), colour_by = "p", text_by = "p") |>
pan_camera("right lateral")Hover over the coloured regions — the tooltip now shows each region’s p-value.
Custom colour palettes
An unnamed vector defines the gradient endpoints:
ggseg3d(
.data = some_data,
atlas = dk(),
colour_by = "p",
palette = c("forestgreen", "white", "firebrick")
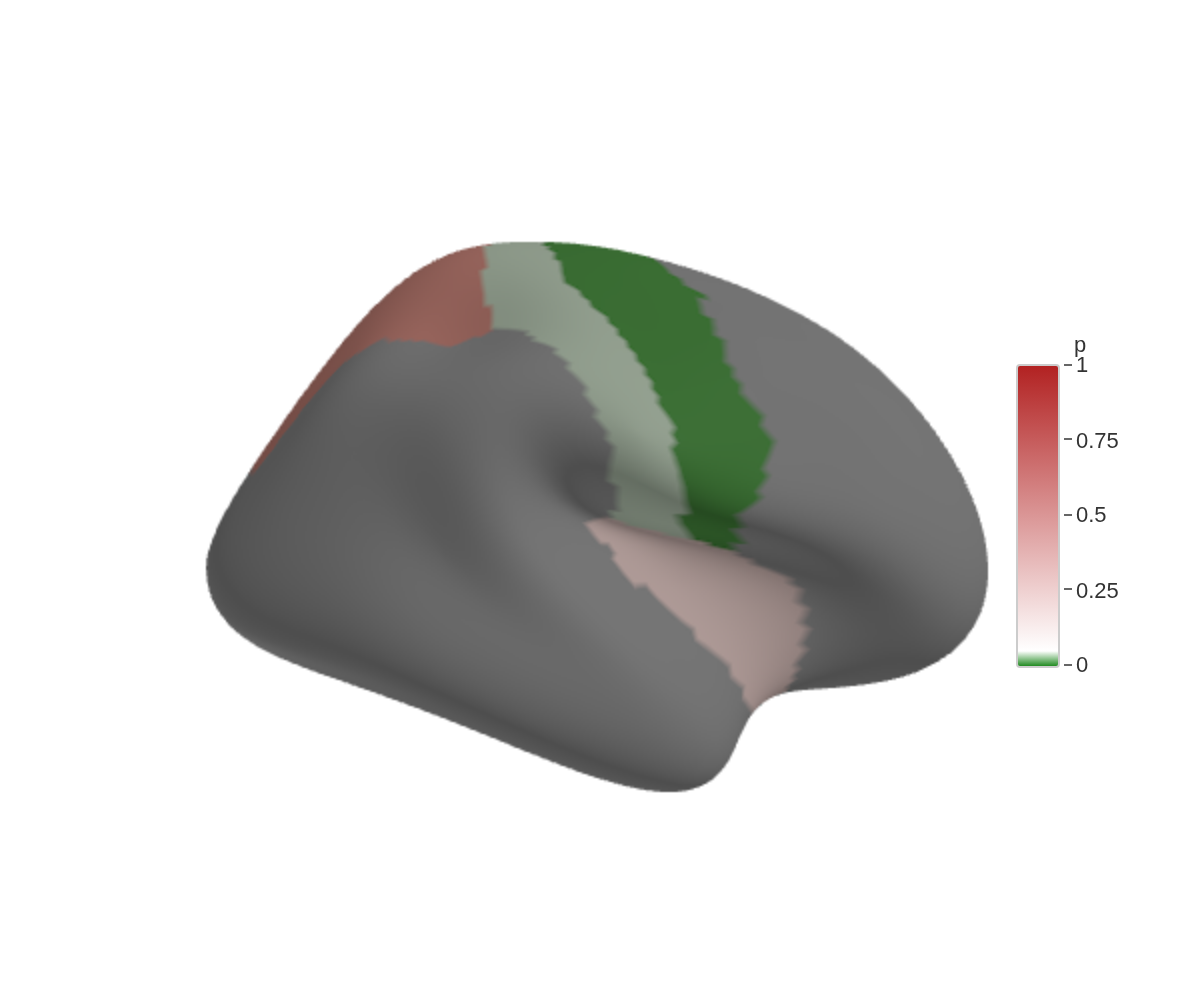
)A named vector pins colours to specific breakpoints, which lets the scale extend beyond your data range:
ggseg3d(
.data = some_data,
atlas = dk(),
colour_by = "p",
text_by = "p",
palette = c("forestgreen" = 0, "white" = .05, "firebrick" = 1)
)
Background colour
White works for most journals. Dark backgrounds work for slides:
ggseg3d() |>
set_background("black")
Region edges
Experimental. Edge rendering works in the htmlwidget viewer. rgl support (
ggsegray) is still unreliable across platforms.
set_edges() draws outlines along region boundaries:
ggseg3d(hemisphere = "left") |>
set_edges("black", width = 10) |>
pan_camera("left lateral")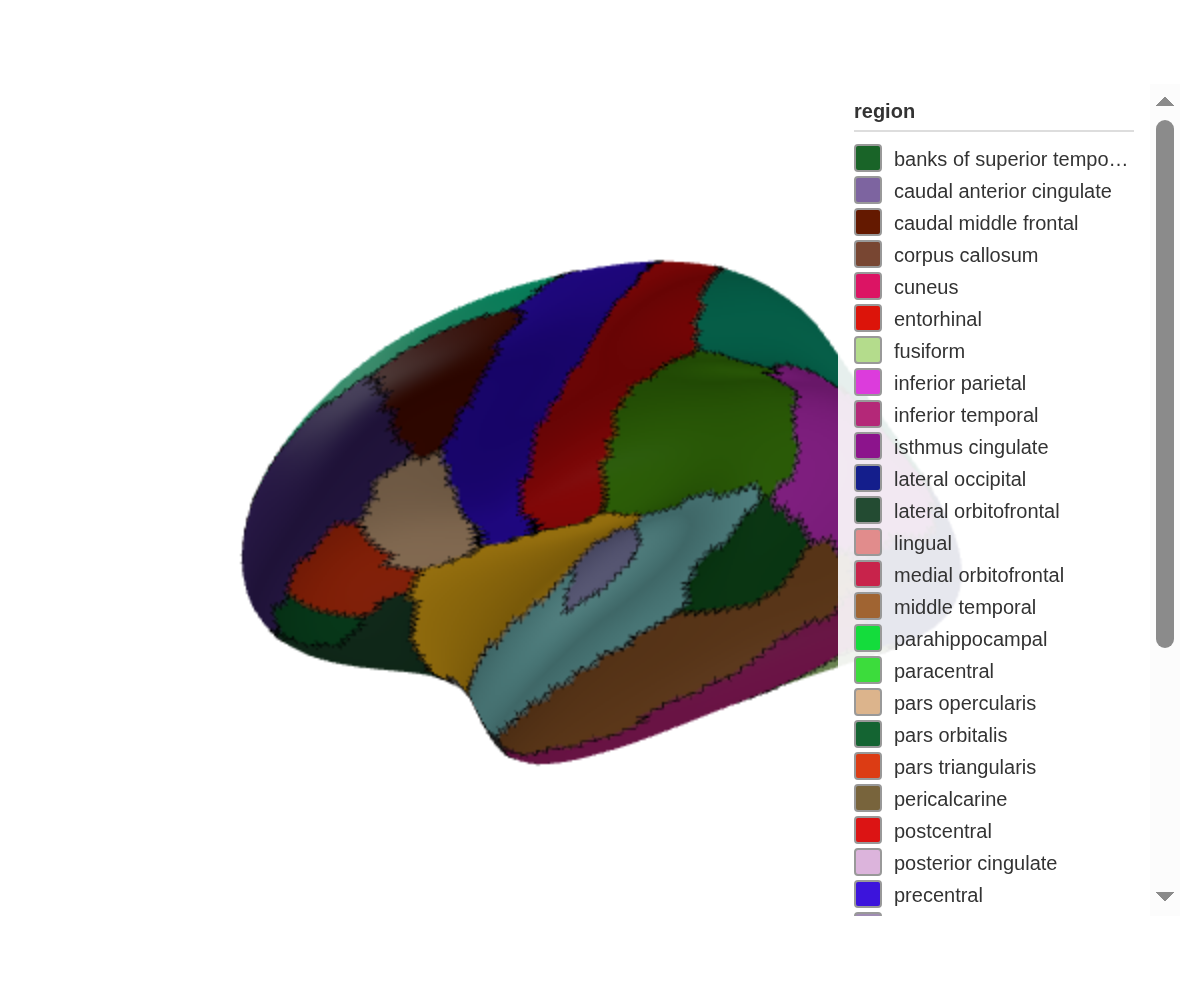
Grouping edges with edge_by
By default, edges appear wherever adjacent vertices have different
colours. edge_by lets you group edges by a different column
— useful when you want boundaries between lobes but not between
individual regions within the same lobe:
lobe_data <- tibble(
region = c("precentral", "postcentral", "insula", "superior parietal"),
p = c(0.2, 0.4, 0.3, 0.5),
lobe = c("frontal", "parietal", "insula", "parietal")
)
ggseg3d(.data = lobe_data, atlas = dk(), colour_by = "p", edge_by = "lobe") |>
set_edges("white", width = 1) |>
set_background("black")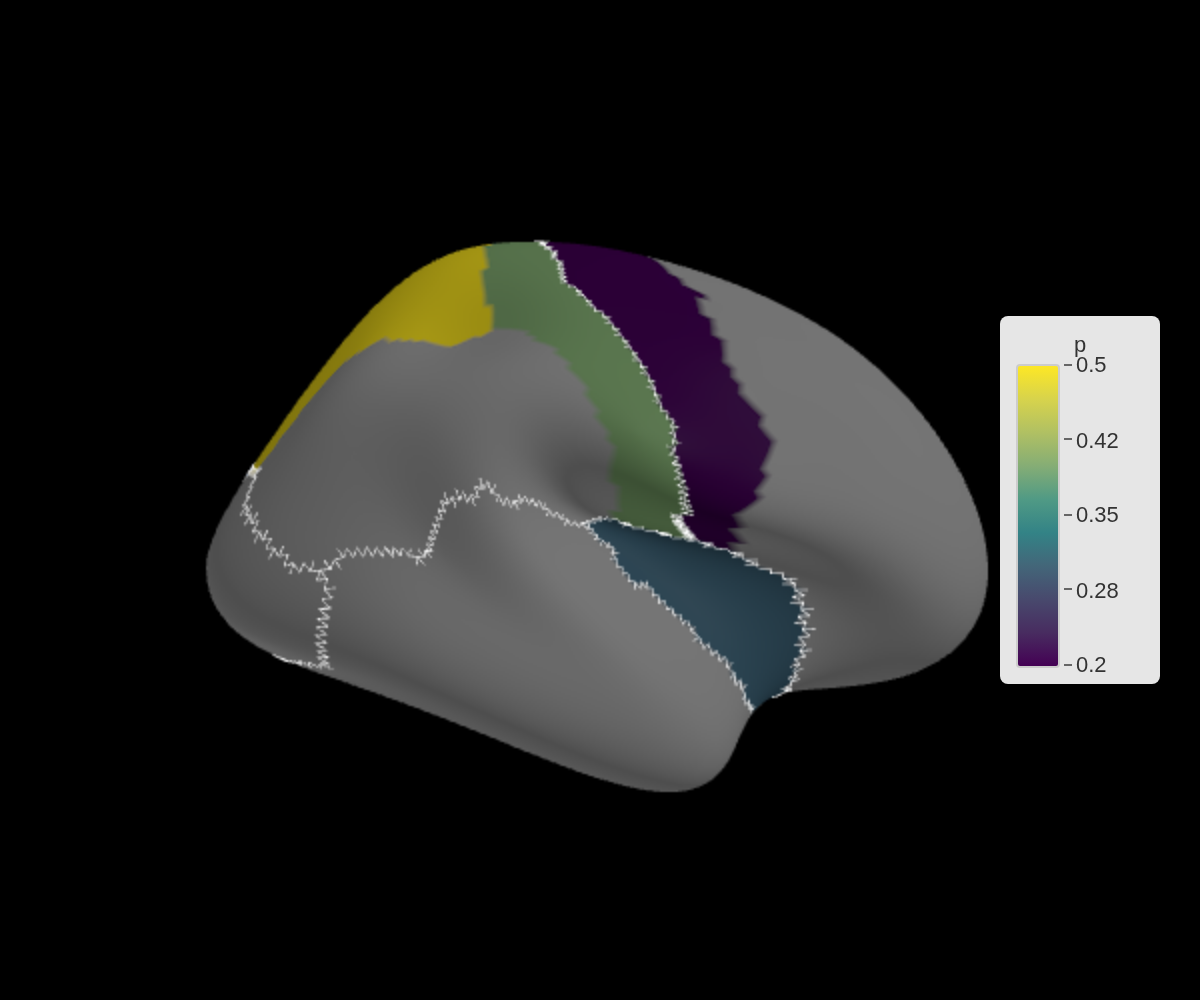
Subcortical structures
The aseg atlas covers structures beneath the cortex.
Fade unmatched regions with na_alpha and wrap everything in
a glass brain:
subcort_data <- tibble(
region = c("Thalamus", "Caudate", "Hippocampus"),
p = c(0.2, 0.5, 0.8)
)
ggseg3d(
.data = subcort_data,
atlas = aseg(),
colour_by = "p",
text_by = "p",
na_alpha = .5
) |>
add_glassbrain()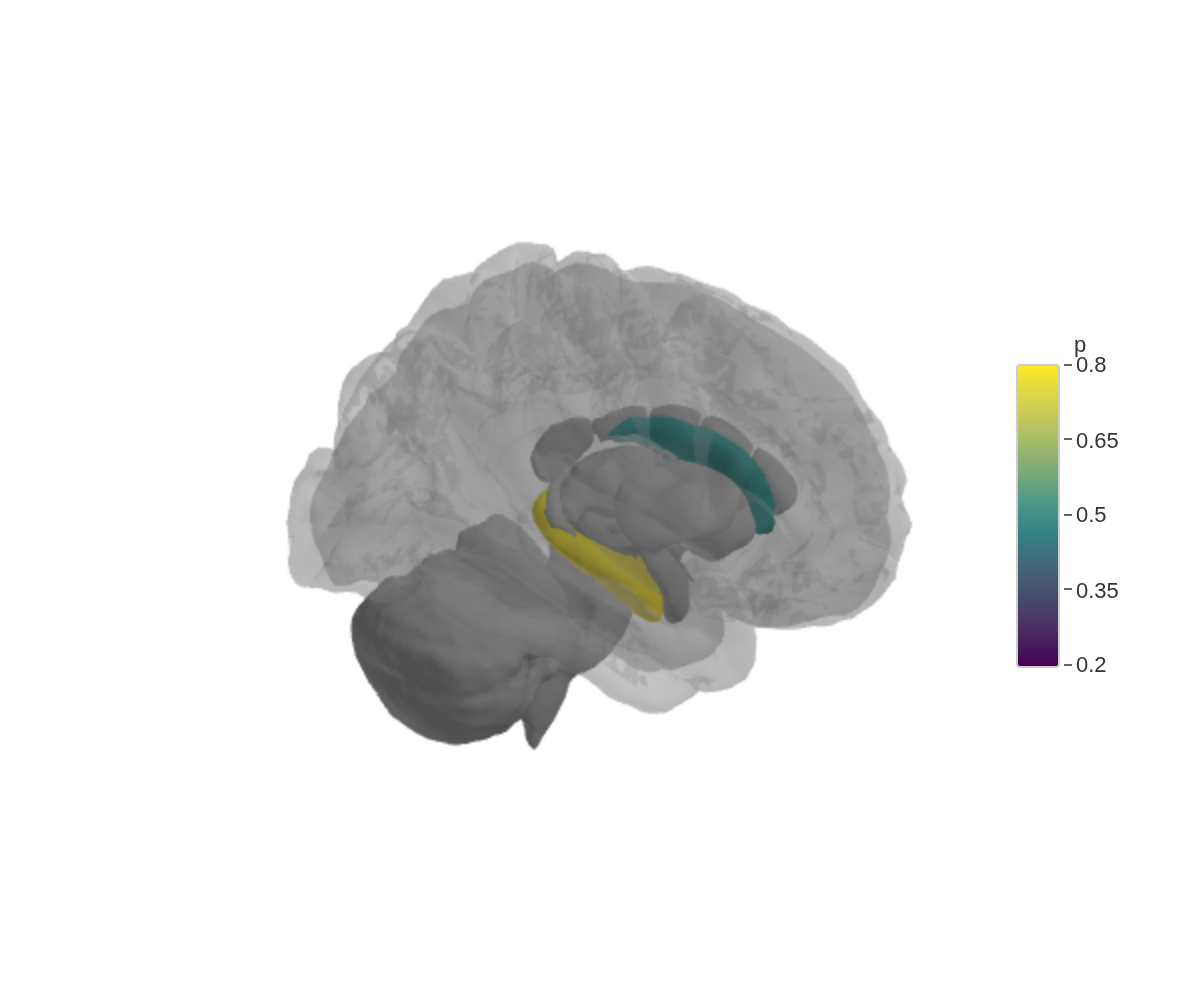
The glass brain is just a translucent cortical surface — enough anatomy to orient the viewer without hiding the structures underneath.
White matter tracts
The tracula atlas renders white matter pathways as tube
meshes. A near-invisible glass brain keeps things oriented:
ggseg3d(atlas = tracula()) |>
add_glassbrain(opacity = 0.1)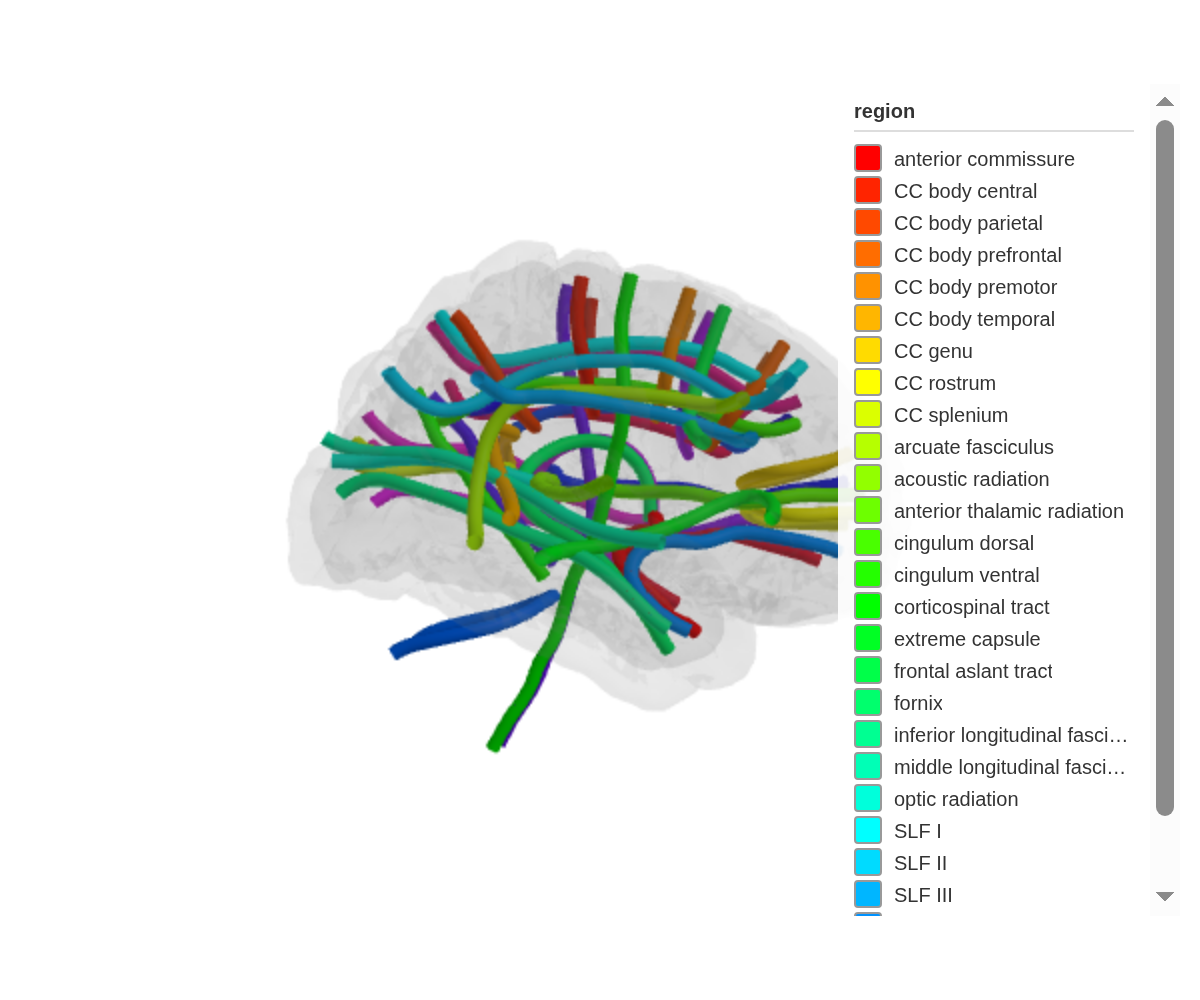
Mapping data to tracts
Data mapping works the same way as cortical and subcortical atlases —
the region column matches tract names:
tract_data <- tibble(
region = c("arcuate fasciculus", "corticospinal tract",
"cingulum bundle", "uncinate fasciculus"),
fa = c(0.45, 0.55, 0.40, 0.35)
)
ggseg3d(
.data = tract_data,
atlas = tracula(),
colour_by = "fa",
text_by = "fa"
) |>
add_glassbrain(opacity = 0.1)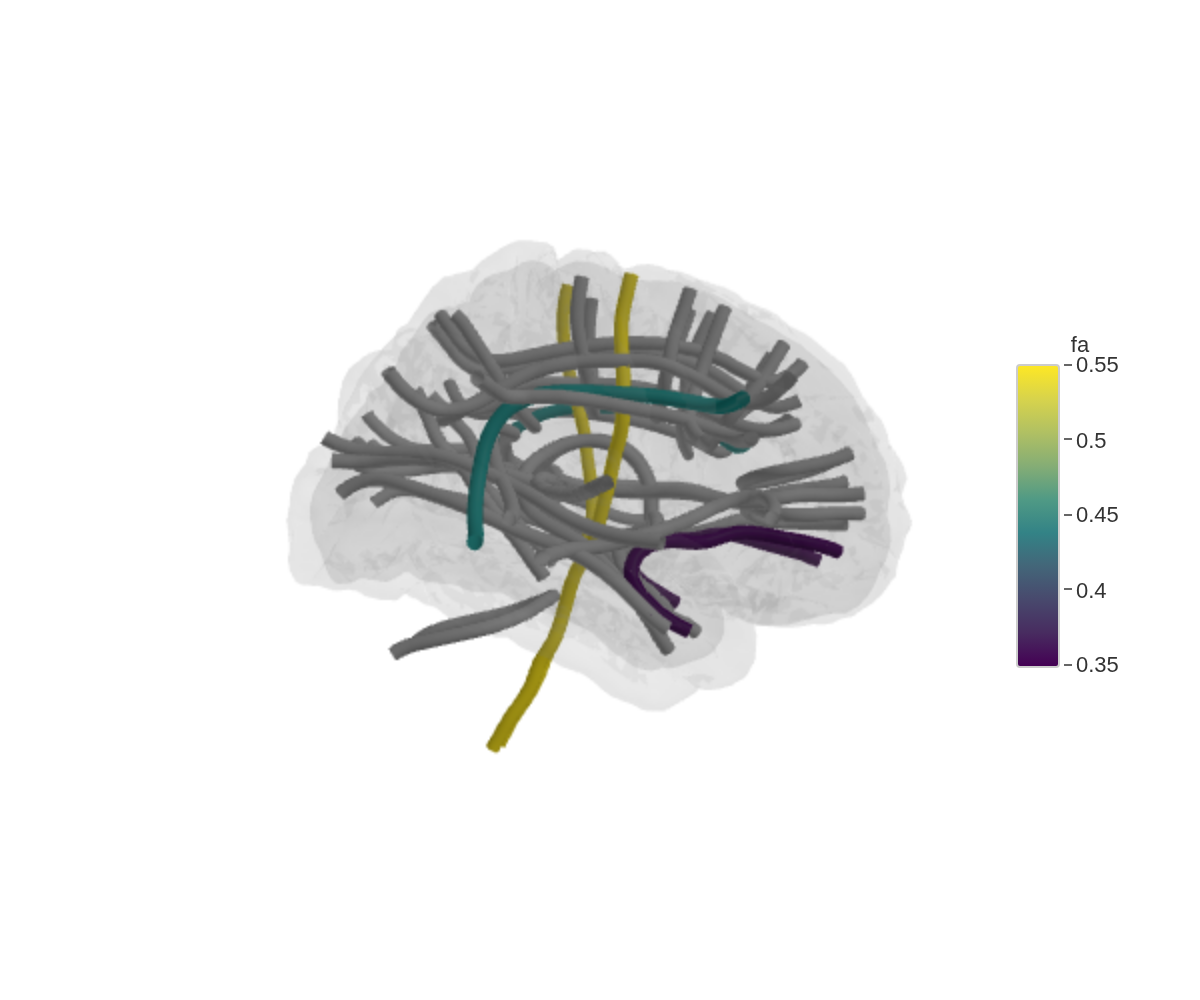
Orientation colouring
tract_color = "orientation" encodes fibre direction as
RGB — red for left-right, green for anterior-posterior, blue for
superior-inferior. This is the same convention used in DTI colour
maps:
ggseg3d(atlas = tracula(), tract_color = "orientation") |>
add_glassbrain(opacity = 0.1) |>
set_background("black")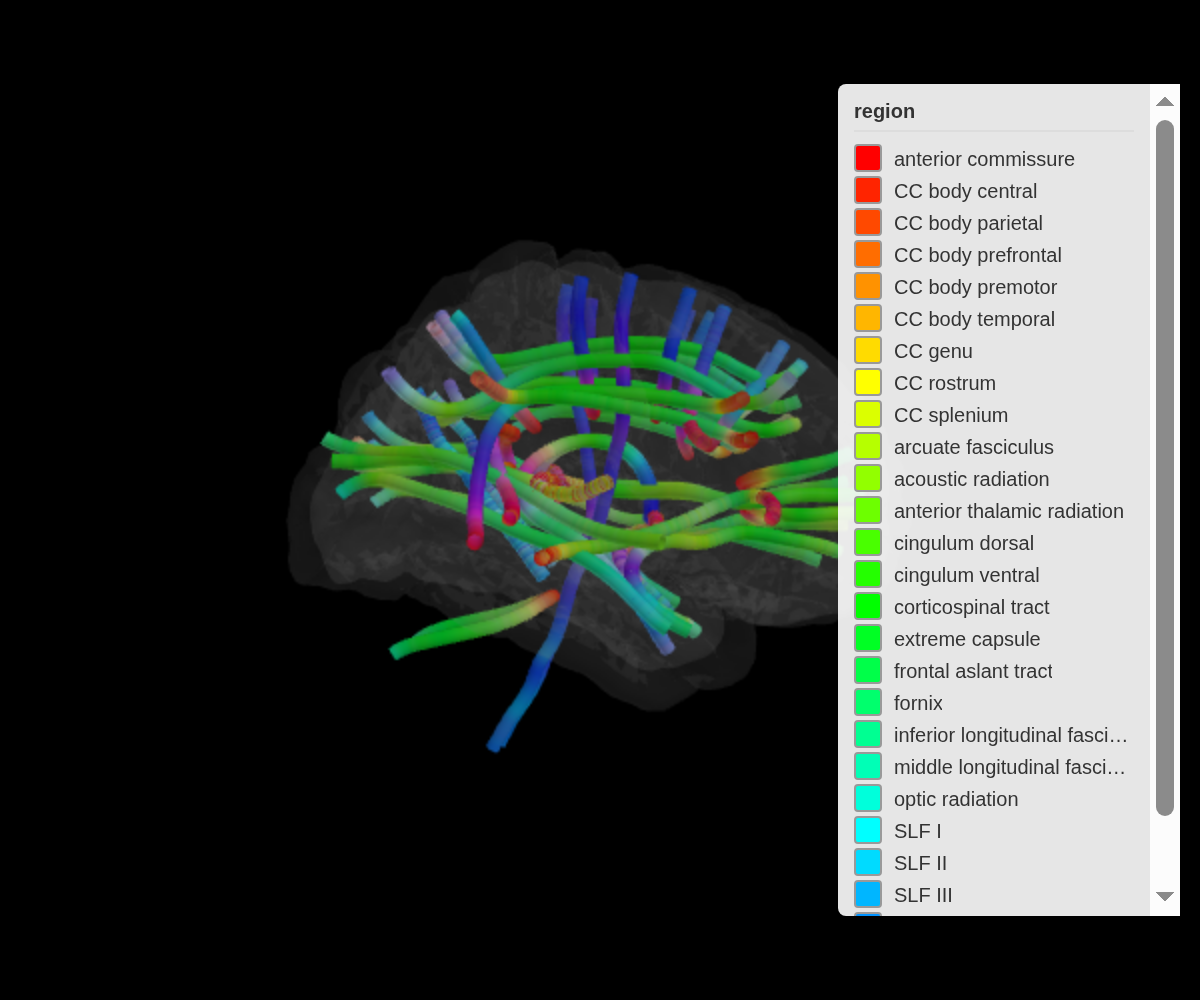
Tube appearance
tube_radius controls thickness,
tube_segments controls smoothness. Higher segments look
better but render slower:
ggseg3d(atlas = tracula(), tube_radius = 4, tube_segments = 16) |>
add_glassbrain(opacity = 0.1)Saving images
snapshot_brain() renders the widget to a PNG using a
headless browser (Chrome or Chromium required):
ggseg3d() |>
pan_camera("left lateral") |>
snapshot_brain("brain_lateral.png")Publication-quality settings
For print, crank up the resolution:
ggseg3d(.data = some_data, atlas = dk(), colour_by = "p") |>
pan_camera("left lateral") |>
set_background("white") |>
snapshot_brain(
"figure1_brain.png",
width = 1200,
height = 1000,
zoom = 3,
delay = 2
)- width/height — output dimensions in pixels
- zoom — resolution multiplier (2-3 for print)
- delay — seconds to wait for the renderer to finish
Multi-panel figures
Snapshot each view separately, then stitch them together with magick:
library(magick)
base_plot <- ggseg3d(.data = some_data, atlas = dk(), colour_by = "p") |>
set_background("white")
base_plot |>
pan_camera("left lateral") |>
snapshot_brain("left_lat.png")
base_plot |>
pan_camera("left medial") |>
snapshot_brain("left_med.png")
base_plot |>
pan_camera("right lateral") |>
snapshot_brain("right_lat.png")
base_plot |>
pan_camera("right medial") |>
snapshot_brain("right_med.png")
panels <- image_read(c(
"left_lat.png",
"left_med.png",
"right_lat.png",
"right_med.png"
))
combined <- image_montage(panels, geometry = "600x500", tile = "2x2")
image_write(combined, "figure1_all_views.png")## agg_png
## 2## [1] TRUE TRUE TRUE TRUE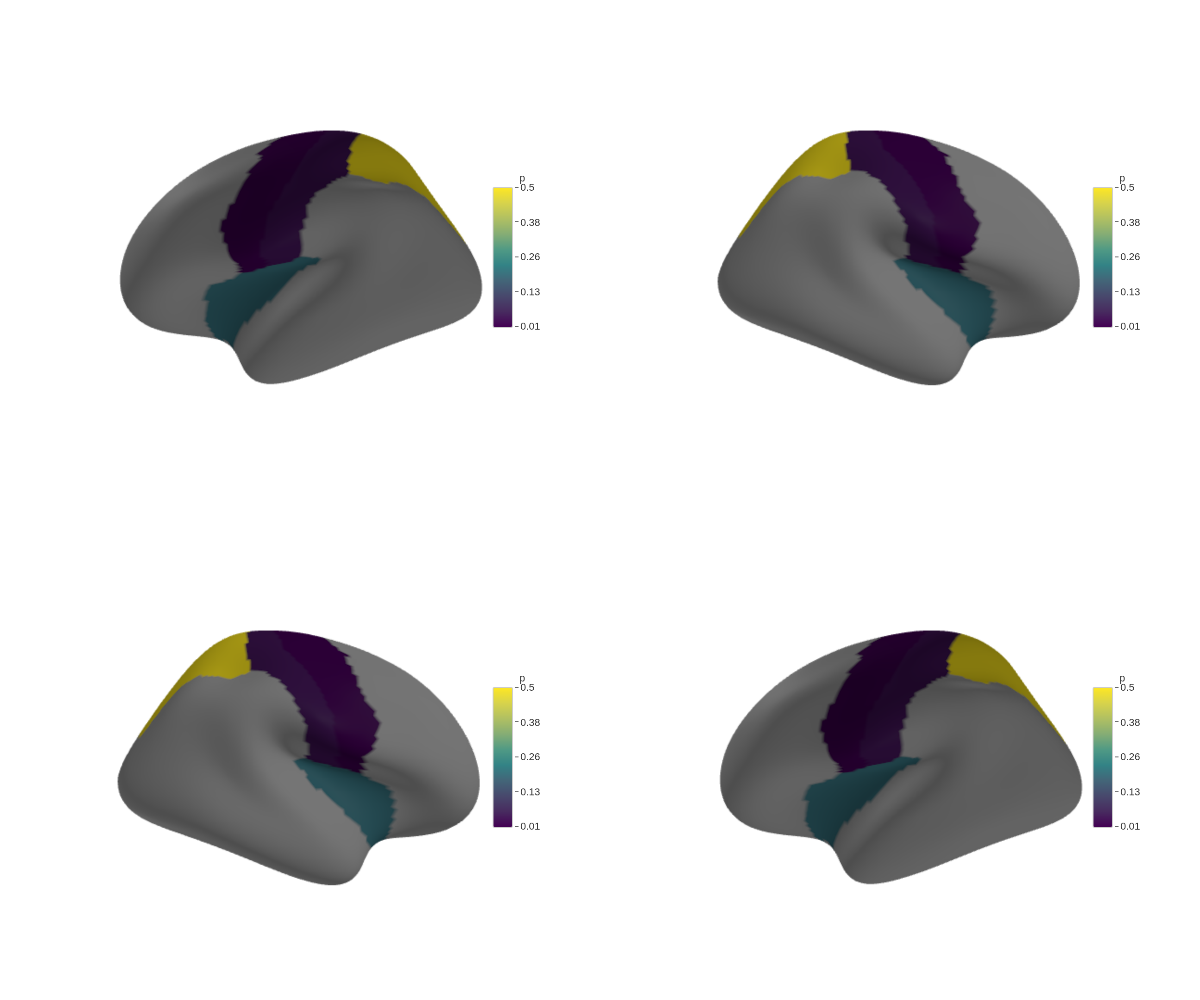
Legend control
ggseg3d() |>
set_legend(FALSE)Widget dimensions
ggseg3d() |>
set_dimensions(width = 800, height = 600)Flat shading
Lighting adds depth but shifts colours.
set_flat_shading() turns off all lighting so every vertex
renders at its exact assigned colour — useful for atlas illustrations or
mask extraction:
ggseg3d() |>
set_flat_shading()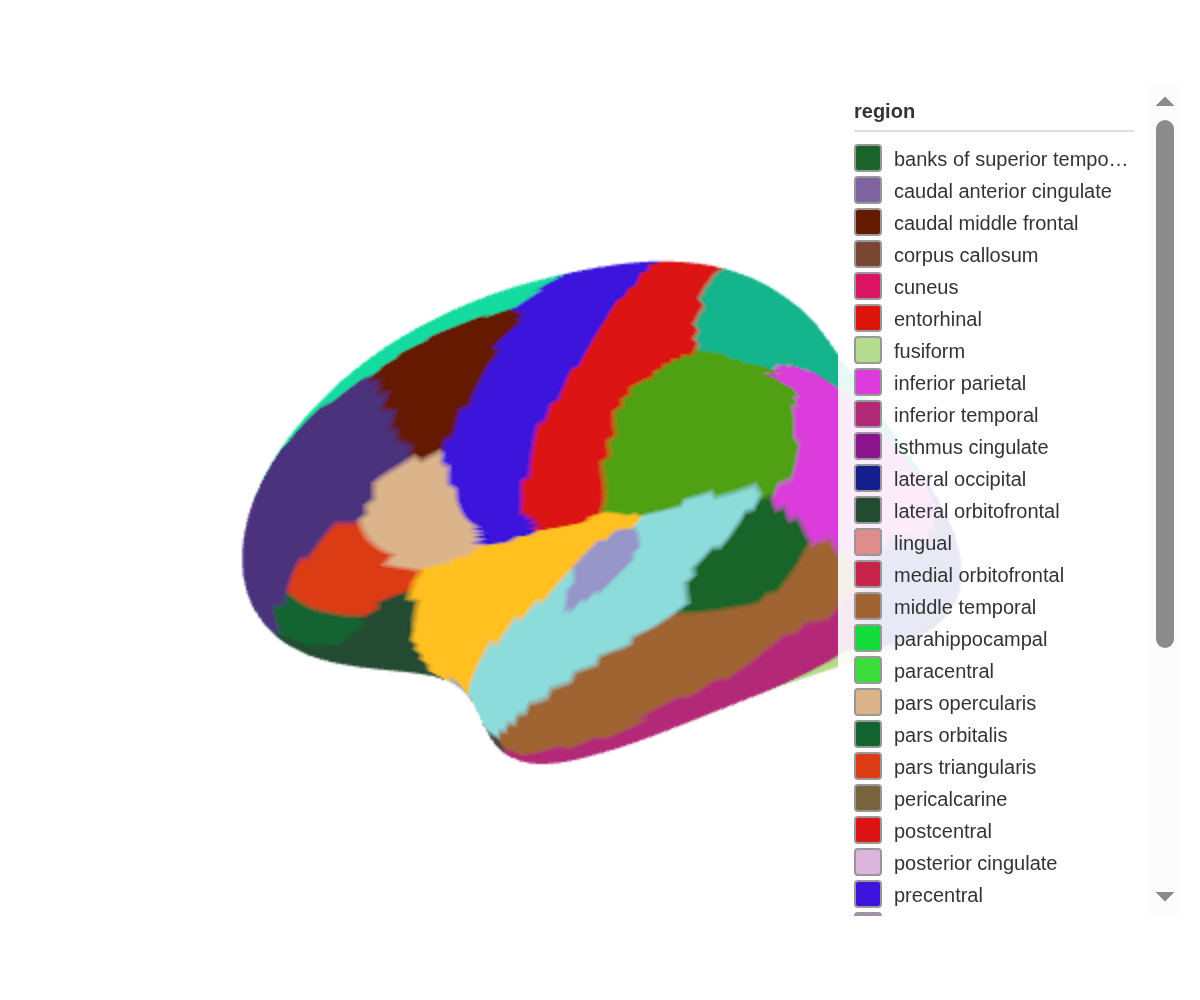
Orthographic projection
Perspective projection makes distant regions look smaller. Orthographic projection removes that depth cue, so every region appears at the same scale regardless of distance from the camera:
ggseg3d() |>
set_orthographic()