from plotnine import ggplot
from ggsegpy import geom_brain, dk
ggplot() + geom_brain(atlas=dk())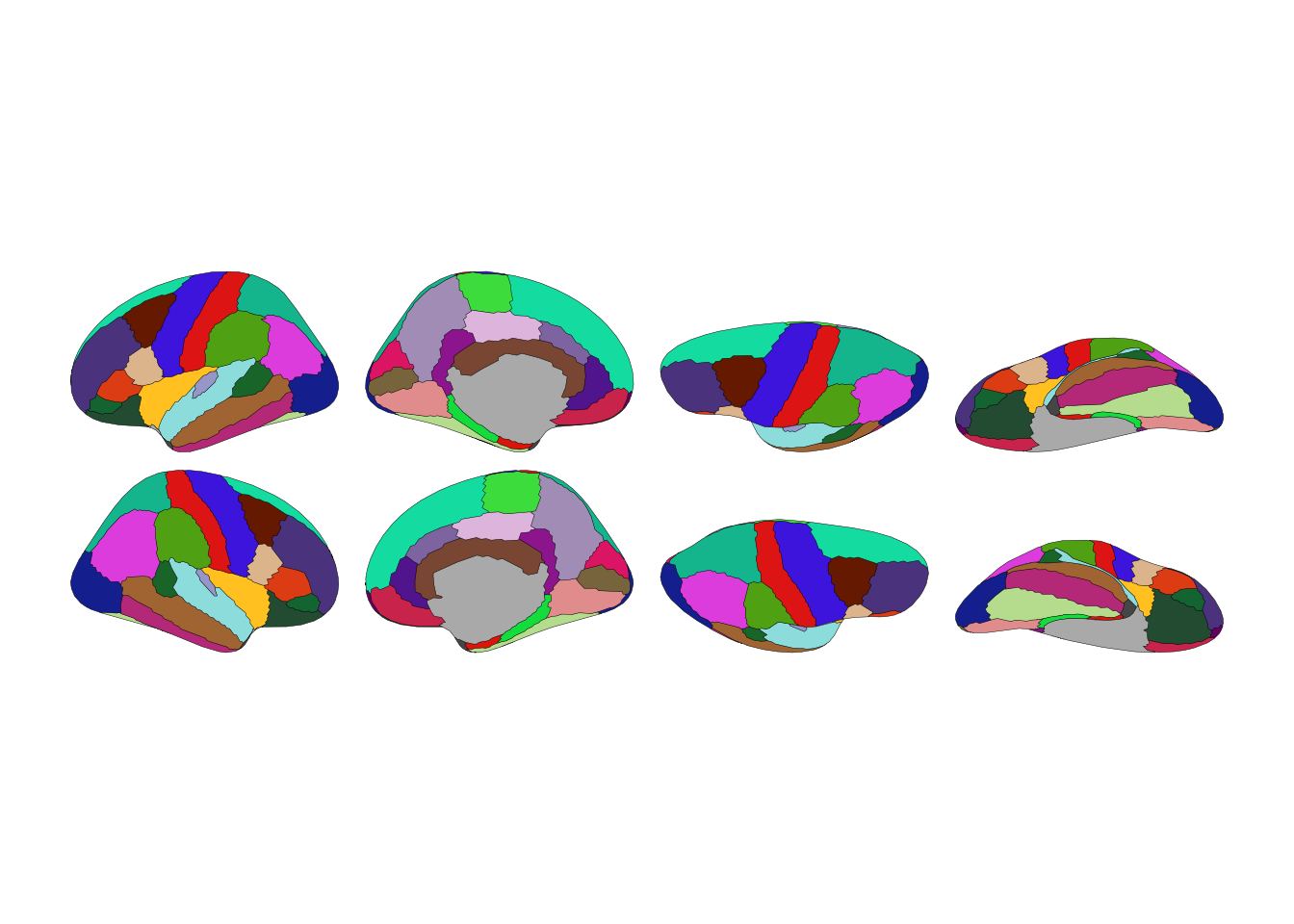
Brain atlas visualization in Python
Three lines to visualize a brain atlas:
ggsegpy is a Python port of the R ggseg ecosystem. It provides 2D visualization with plotnine and 3D interactive visualization with Plotly.
Three atlases ship with the package:
| Atlas | Type | Regions |
|---|---|---|
dk() |
Cortical | Desikan-Killiany parcellation (34 × 2 hemispheres) |
aseg() |
Subcortical | FreeSurfer subcortical segmentation |
tracula() |
White matter | TRACULA tract atlas |
Each works in both 2D and 3D: