Sometimes you have brain regions defined as individual FreeSurfer label files rather than a complete annotation. This happens when you extract results from vertex-wise analyses, combine regions from different sources, or work with manually defined ROIs.
create_cortical_from_labels() builds an atlas from a collection of .label files.
What label files contain
FreeSurfer label files are text files that list which vertices belong to a region:
#!ascii label
5432
1234 -23.5 12.3 45.6 0.0
1235 -23.8 12.1 45.9 0.0
...The first line is a header, the second is the vertex count, and subsequent lines contain the vertex index and its coordinates. Label files conventionally include hemisphere in the filename: lh.motor.label or rh.motor.label.
Basic usage
Collect your label files and create an atlas:
label_dir <- file.path(
freesurfer::fs_dir(), "subjects", "fsaverage5", "label"
)
ba_labels <- list.files(
label_dir,
pattern = "^[lr]h\\.BA[0-9]+.*\\.label$",
full.names = TRUE
)
length(ba_labels)
#> [1] 18
head(basename(ba_labels))
#> [1] "lh.BA1_exvivo.label"
#> [2] "lh.BA2_exvivo.label"
#> [3] "lh.BA3a_exvivo.label"
#> [4] "lh.BA3b_exvivo.label"
#> [5] "lh.BA44_exvivo.label"
#> [6] "lh.BA45_exvivo.label"
ba_atlas <- create_cortical_from_labels(
label_files = ba_labels,
atlas_name = "brodmann"
)
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef86163473491.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef861263665d6.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef86152331bbb.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef8615f876fce.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef86199bd54.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef86169ec7d0d.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef86132c38306.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef86136f1b46.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef86114be2ba5.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef8616b92f547.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef861186b67bb.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef8613c2550d2.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef8614ae5fde1.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef86171f9158e.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef8612e115a4b.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef8617d8c441d.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef861493c5a22.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef861204aabe6.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef86167c150b8.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef861bc7e49.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef86172e37fc.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef86148cb97cf.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef8618ac18c0.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef8615942d38f.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef86111f4ee5.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef86169ecf24d.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef861514273b2.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef8616f418d8c.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef86138725f51.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef8615266da60.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef86177641992.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef86122c77bf9.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef861269b446f.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef8612f8a986a.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef86136de6472.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef86128a204f4.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef8615ef633e8.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef8611df7233e.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef861d6944c1.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef8614c70e75.html screenshot completed
ba_atlas
#>
#> ── brodmann ggseg atlas ──────────────────
#> Type: cortical
#> Regions: 9
#> Hemispheres: left, right
#> Views: lateral, medial
#> Palette: ✖
#> Rendering: ✔ ggseg
#> ✔ ggseg3d (vertices)
#> ──────────────────────────────────────────
#> # A tibble: 18 × 3
#> hemi region label
#> <chr> <chr> <chr>
#> 1 left BA1_exvivo lh_BA1_exvivo
#> 2 left BA2_exvivo lh_BA2_exvivo
#> 3 left BA3a_exvivo lh_BA3a_exvivo
#> 4 left BA3b_exvivo lh_BA3b_exvivo
#> 5 left BA44_exvivo lh_BA44_exvivo
#> 6 left BA45_exvivo lh_BA45_exvivo
#> 7 left BA4a_exvivo lh_BA4a_exvivo
#> 8 left BA4p_exvivo lh_BA4p_exvivo
#> 9 left BA6_exvivo lh_BA6_exvivo
#> 10 right BA1_exvivo rh_BA1_exvivo
#> 11 right BA2_exvivo rh_BA2_exvivo
#> 12 right BA3a_exvivo rh_BA3a_exvivo
#> 13 right BA3b_exvivo rh_BA3b_exvivo
#> 14 right BA44_exvivo rh_BA44_exvivo
#> 15 right BA45_exvivo rh_BA45_exvivo
#> 16 right BA4a_exvivo rh_BA4a_exvivo
#> 17 right BA4p_exvivo rh_BA4p_exvivo
#> 18 right BA6_exvivo rh_BA6_exvivoThe function detects hemisphere from filename prefixes (lh. or rh.) and derives region names from the rest of the filename.
Custom names and colours
Override auto-detected names when the filenames don’t produce readable labels:
atlas <- create_cortical_from_labels(
label_files = c(
"lh.motor.label", "rh.motor.label",
"lh.visual.label", "rh.visual.label"
),
region_names = c(
"primary motor", "primary motor",
"primary visual", "primary visual"
),
colours = c("#E74C3C", "#E74C3C", "#3498DB", "#3498DB")
)Left and right hemisphere versions of the same region should share the same region name — the hemisphere column distinguishes them.
Full pipeline with 2D geometry
For 2D plots, enable the full pipeline:
ba_atlas <- create_cortical_from_labels(
label_files = ba_labels,
atlas_name = "custom"
)
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef86110f02b1a.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef8615d2d632.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef8616e931304.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef8611ea741f3.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef861159059c1.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef8611b665b2d.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef86173cae81.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef86173340a.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef86126a968bb.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef86189dd82b.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef86144a6655d.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef86168623106.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef8616c4a7d01.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef8616f8675f7.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef861f747a6e.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef8612a609ab6.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef8611b5e1b3.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef8612d37a098.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef86170c340fd.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef86116dca2aa.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef8611fad7dc6.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef8612880f7c3.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef8612be155d0.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef8611e38847a.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef861232dffd1.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef8617799ced5.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef86136715e98.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef861a61c787.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef86129790549.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef861769b0dcf.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef86153bcc971.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef86119d6fcbf.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef86143b754bb.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef86156036252.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef861195284c0.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef8614972af7.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef8616d1ffcbe.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef8612f8e9e30.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef861577b374c.html screenshot completed
#> file:////private/var/folders/y5/zlbbcqn56gx1tcg6xfb425100000gp/T/RtmpTTP4iV/filef8614a9017be.html screenshot completed
ba_atlas
#>
#> ── custom ggseg atlas ────────────────────
#> Type: cortical
#> Regions: 9
#> Hemispheres: left, right
#> Views: lateral, medial
#> Palette: ✖
#> Rendering: ✔ ggseg
#> ✔ ggseg3d (vertices)
#> ──────────────────────────────────────────
#> # A tibble: 18 × 3
#> hemi region label
#> <chr> <chr> <chr>
#> 1 left BA1_exvivo lh_BA1_exvivo
#> 2 left BA2_exvivo lh_BA2_exvivo
#> 3 left BA3a_exvivo lh_BA3a_exvivo
#> 4 left BA3b_exvivo lh_BA3b_exvivo
#> 5 left BA44_exvivo lh_BA44_exvivo
#> 6 left BA45_exvivo lh_BA45_exvivo
#> 7 left BA4a_exvivo lh_BA4a_exvivo
#> 8 left BA4p_exvivo lh_BA4p_exvivo
#> 9 left BA6_exvivo lh_BA6_exvivo
#> 10 right BA1_exvivo rh_BA1_exvivo
#> 11 right BA2_exvivo rh_BA2_exvivo
#> 12 right BA3a_exvivo rh_BA3a_exvivo
#> 13 right BA3b_exvivo rh_BA3b_exvivo
#> 14 right BA44_exvivo rh_BA44_exvivo
#> 15 right BA45_exvivo rh_BA45_exvivo
#> 16 right BA4a_exvivo rh_BA4a_exvivo
#> 17 right BA4p_exvivo rh_BA4p_exvivo
#> 18 right BA6_exvivo rh_BA6_exvivoThis runs the same screenshot pipeline as create_cortical_from_annotation() — it takes screenshots of each region from multiple angles, extracts contours, and converts them to sf polygons.
Post-processing
Clean up region names:
ba_atlas <- ba_atlas |>
atlas_region_contextual("cortex", match_on = "label") |>
atlas_region_contextual("unknown", match_on = "label") |>
atlas_view_gather()
core_clean <- ba_atlas$core |>
mutate(
region = gsub("\\.", " ", region),
region = gsub("_exvivo", "", region)
)
ba_atlas <- ggseg_atlas(
atlas = ba_atlas$atlas,
type = ba_atlas$type,
palette = ba_atlas$palette,
core = core_clean,
data = ba_atlas$data
)
plot(ba_atlas) +
ggplot2::scale_fill_viridis_d(na.value = "grey80")
#> Scale for fill is already present.
#> Adding another scale for fill, which will
#> replace the existing scale.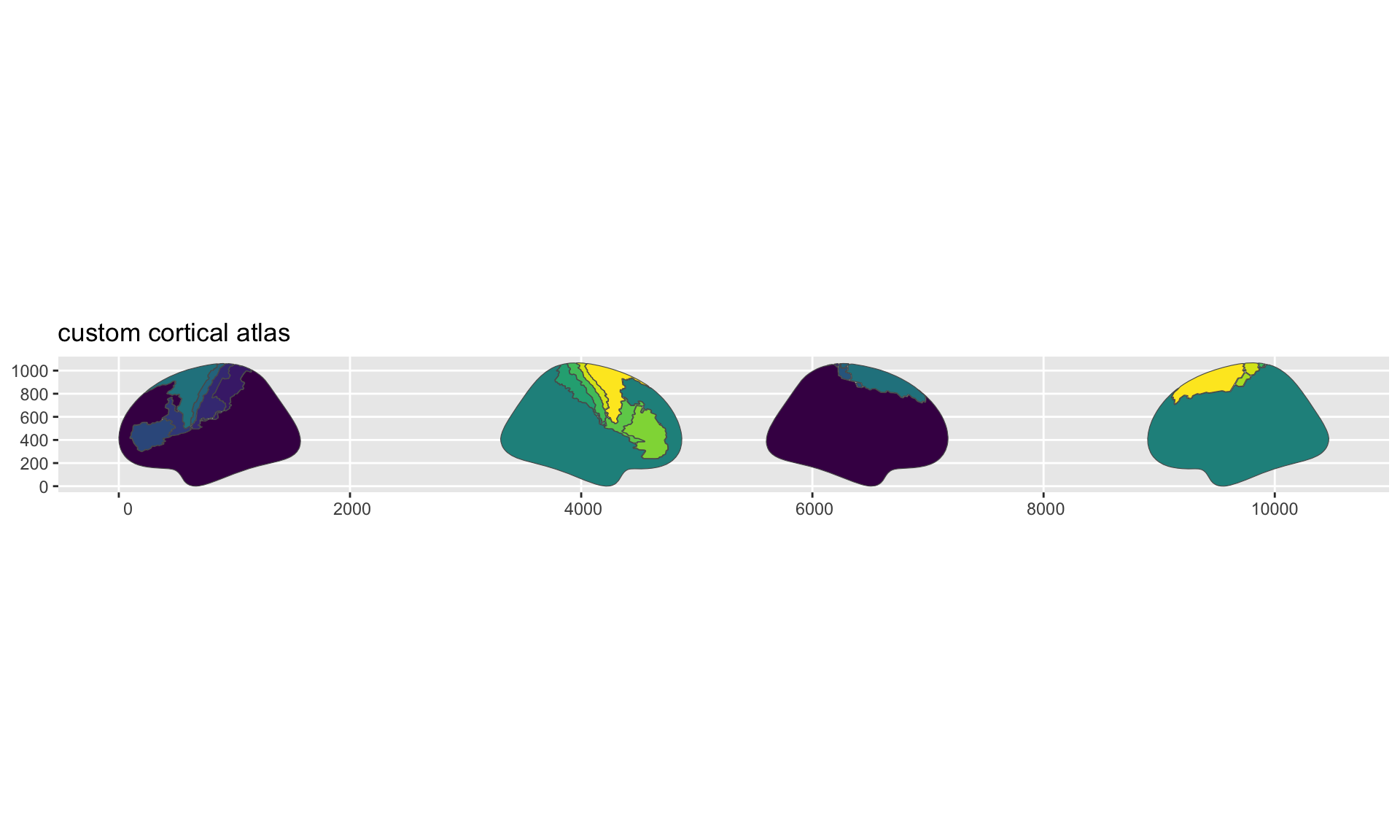
Brodmann area atlas plotted with ggseg.
Finding label files
FreeSurfer ships with several sets of label files. Brodmann areas are the most common:
label_dir <- file.path(
freesurfer::fs_dir(), "subjects", "fsaverage5", "label"
)
ba_files <- list.files(label_dir, "^[lr]h\\.BA.*\\.label$", full.names = TRUE)
length(ba_files)
#> [1] 18Other useful labels include V1/V2, MT, and entorhinal cortex. Any label file that maps to the same surface (typically fsaverage5) can be combined into one atlas.
Combining regions from different sources
Label-based atlases are useful when you want to mix regions from different analyses or atlases. Each label file is independent, so you can combine freely:
my_labels <- c(
"lh.BA1_exvivo.label",
"lh.BA2_exvivo.label",
"lh.V1.label",
"rh.BA1_exvivo.label",
"rh.BA2_exvivo.label",
"rh.V1.label"
)
atlas <- create_cortical_from_labels(
label_files = file.path(label_dir, my_labels),
atlas_name = "somatosensory_visual",
region_names = c(
"BA1", "BA2", "V1",
"BA1", "BA2", "V1"
),
colours = c(
"#E74C3C", "#C0392B", "#3498DB",
"#E74C3C", "#C0392B", "#3498DB"
)
)The only constraint is that all label files must reference the same surface mesh. Labels created for fsaverage won’t match fsaverage5 vertices without resampling.