Tract atlases represent white matter pathways derived from diffusion MRI tractography. Unlike cortical or subcortical atlases where regions are contiguous surfaces, tracts are tube-like structures that follow streamline bundles through the brain.
The pipeline reads tractography files, computes a centerline for each tract, wraps it in a tube mesh, and (optionally) creates 2D projection views.
This tutorial recreates the TRACULA atlas — the same pipeline behind data-raw/make_tracula_atlas.R in ggseg.formats.
What you need
- FreeSurfer installed with TRACULA training data (
trctrain/) - ImageMagick for 2D geometry extraction
- Chrome/Chromium for 3D screenshots
Finding tractography files
TRACULA training data ships with FreeSurfer in the trctrain/ directory. Each .trk file contains streamlines for one tract:
fs_dir <- freesurfer::fs_dir()
tract_dir <- file.path(fs_dir, "trctrain", "hcp", "mgh_1017", "mni")
tract_files <- list.files(tract_dir, pattern = "\\.trk$", full.names = TRUE)
aseg_file <- file.path(tract_dir, "aparc+aseg.nii.gz")
length(tract_files)
#> [1] 42The aparc+aseg.nii.gz provides cortex outlines for 2D projections. If it’s missing, you can still create a 3D-only atlas.
Creating the atlas
create_tract_from_tractography() reads each tractography file, computes a centerline, and generates a tube mesh around it. The input_aseg provides cortex context for 2D views:
tracula_raw <- create_tract_from_tractography(
input_tracts = tract_files,
input_aseg = aseg_file,
atlas_name = "tracula"
)
#> Warning: Atlas has 11230 vertices (threshold:
#> 10000)
#> ℹ Large atlases may be slow to plot and
#> increase package size
#> ℹ Re-run with higher `tolerance` to
#> reduce vertices
tracula_raw
#>
#> ── tracts ggseg atlas ────────────────────
#> Type: tract
#> Regions: 26
#> Hemispheres: midline, left, right
#> Views: axial_1, axial_2, axial_3,
#> axial_4, axial_5, coronal_1, coronal_2,
#> coronal_3, coronal_4, coronal_5,
#> coronal_6, sagittal_left,
#> sagittal_midline, sagittal_right
#> Palette: ✖
#> Rendering: ✔ ggseg
#> ✔ ggseg3d (centerlines)
#> ──────────────────────────────────────────
#> # A tibble: 42 × 3
#> hemi region label
#> <chr> <chr> <chr>
#> 1 midline acomm.bbr.prep acomm.bbr…
#> 2 midline cc.bodyc.bbr.prep cc.bodyc.…
#> 3 midline cc.bodyp.bbr.prep cc.bodyp.…
#> 4 midline cc.bodypf.bbr.prep cc.bodypf…
#> 5 midline cc.bodypm.bbr.prep cc.bodypm…
#> 6 midline cc.bodyt.bbr.prep cc.bodyt.…
#> 7 midline cc.genu.bbr.prep cc.genu.b…
#> 8 midline cc.rostrum.bbr.prep cc.rostru…
#> 9 midline cc.splenium.bbr.prep cc.spleni…
#> 10 left af.bbr.prep lh.af.bbr…
#> 11 left ar.bbr.prep lh.ar.bbr…
#> 12 left atr.bbr.prep lh.atr.bb…
#> 13 left cbd.bbr.prep lh.cbd.bb…
#> 14 left cbv.bbr.prep lh.cbv.bb…
#> 15 left cst.bbr.prep lh.cst.bb…
#> 16 left emc.bbr.prep lh.emc.bb…
#> 17 left fat.bbr.prep lh.fat.bb…
#> 18 left fx.bbr.prep lh.fx.bbr…
#> 19 left ilf.bbr.prep lh.ilf.bb…
#> 20 left mlf.bbr.prep lh.mlf.bb…
#> 21 left or.bbr.prep lh.or.bbr…
#> 22 left slf1.bbr.prep lh.slf1.b…
#> 23 left slf2.bbr.prep lh.slf2.b…
#> 24 left slf3.bbr.prep lh.slf3.b…
#> 25 left uf.bbr.prep lh.uf.bbr…
#> 26 midline mcp.bbr.prep mcp.bbr.p…
#> 27 right af.bbr.prep rh.af.bbr…
#> 28 right ar.bbr.prep rh.ar.bbr…
#> 29 right atr.bbr.prep rh.atr.bb…
#> 30 right cbd.bbr.prep rh.cbd.bb…
#> 31 right cbv.bbr.prep rh.cbv.bb…
#> 32 right cst.bbr.prep rh.cst.bb…
#> 33 right emc.bbr.prep rh.emc.bb…
#> 34 right fat.bbr.prep rh.fat.bb…
#> 35 right fx.bbr.prep rh.fx.bbr…
#> 36 right ilf.bbr.prep rh.ilf.bb…
#> 37 right mlf.bbr.prep rh.mlf.bb…
#> 38 right or.bbr.prep rh.or.bbr…
#> 39 right slf1.bbr.prep rh.slf1.b…
#> 40 right slf2.bbr.prep rh.slf2.b…
#> 41 right slf3.bbr.prep rh.slf3.b…
#> 42 right uf.bbr.prep rh.uf.bbr…Key parameters:
-
tube_radiuscontrols tube thickness. Larger values make tracts more visible at the cost of anatomical precision. -
tube_segmentscontrols the number of vertices around the tube circumference. Six gives hexagonal cross-sections; 12+ gives smoother tubes. -
n_pointscontrols centerline resolution. More points capture finer curvature.
Post-processing
Selecting views
Keep the projections that show your tracts best:
tracula_raw <- tracula_raw |>
atlas_view_keep(c(
"axial_2",
"axial_4",
"coronal_3",
"coronal_4",
"sagittal"
))Setting context regions
Add cortex as a background outline:
tracula_raw <- tracula_raw |>
atlas_region_contextual("cortex")Removing small fragments
After view selection, some tracts may appear as tiny slivers in certain views. Remove fragments below a minimum area:
tracula_raw <- tracula_raw |>
atlas_view_remove_small(
min_area = 500,
views = c("axial", "coronal")
) |>
atlas_view_remove_small(min_area = 50)
#> ℹ Removed 49 geometries below area 500
#> ℹ Removed 10 geometries below area 50The first call applies a higher threshold to axial and coronal views (where tracts appear in cross-section and produce small dots). The second call applies a lower threshold everywhere to catch remaining slivers.
Adding metadata
Join tract group information — which tracts are projection fibres, which are commissural, which are association:
normalize_label <- function(x) {
x |>
basename() |>
tools::file_path_sans_ext()
}
tracula_metadata <- data.frame(
label_key = c(
"lh.cst.bbr.prep", "rh.cst.bbr.prep",
"lh.ilf.bbr.prep", "rh.ilf.bbr.prep",
"lh.unc.bbr.prep", "rh.unc.bbr.prep",
"lh.atr.bbr.prep", "rh.atr.bbr.prep",
"lh.cab.bbr.prep", "rh.cab.bbr.prep",
"lh.slfp.bbr.prep", "rh.slfp.bbr.prep",
"lh.slft.bbr.prep", "rh.slft.bbr.prep",
"lh.ccg.bbr.prep", "rh.ccg.bbr.prep",
"fmajor.bbr.prep", "fminor.bbr.prep"
),
region = c(
"corticospinal tract", "corticospinal tract",
"inferior longitudinal fasciculus", "inferior longitudinal fasciculus",
"uncinate fasciculus", "uncinate fasciculus",
"anterior thalamic radiation", "anterior thalamic radiation",
"cingulum (angular bundle)", "cingulum (angular bundle)",
"SLF (parietal)", "SLF (parietal)",
"SLF (temporal)", "SLF (temporal)",
"cingulum (cingulate gyrus)", "cingulum (cingulate gyrus)",
"forceps major", "forceps minor"
),
group = c(
"projection", "projection",
"association", "association",
"association", "association",
"projection", "projection",
"limbic", "limbic",
"association", "association",
"association", "association",
"limbic", "limbic",
"commissural", "commissural"
)
)
core_with_meta <- tracula_raw$core |>
mutate(label_key = normalize_label(label)) |>
left_join(
tracula_metadata |> select(label_key, region_pretty = region, group),
by = "label_key"
) |>
mutate(region = coalesce(region_pretty, region)) |>
select(hemi, region, label, group)Rebuilding and saving
tracula <- ggseg_atlas(
atlas = "tracula",
type = "tract",
palette = tracula_raw$palette,
core = core_with_meta,
data = tracula_raw$data
)
tracula
#>
#> ── tracula ggseg atlas ───────────────────
#> Type: tract
#> Regions: 26
#> Hemispheres: midline, left, right
#> Views: axial_2, axial_4, coronal_3,
#> coronal_4, sagittal_left,
#> sagittal_midline, sagittal_right
#> Palette: ✖
#> Rendering: ✔ ggseg
#> ✔ ggseg3d (centerlines)
#> ──────────────────────────────────────────
#> # A tibble: 42 × 4
#> hemi region label group
#> <chr> <chr> <chr> <chr>
#> 1 midline acomm.bbr.prep acom… <NA>
#> 2 midline cc.bodyc.bbr.prep cc.b… <NA>
#> 3 midline cc.bodyp.bbr.prep cc.b… <NA>
#> 4 midline cc.bodypf.bbr.prep cc.b… <NA>
#> 5 midline cc.bodypm.bbr.prep cc.b… <NA>
#> 6 midline cc.bodyt.bbr.prep cc.b… <NA>
#> 7 midline cc.genu.bbr.prep cc.g… <NA>
#> 8 midline cc.rostrum.bbr.prep cc.r… <NA>
#> 9 midline cc.splenium.bbr.pr… cc.s… <NA>
#> 10 left af.bbr.prep lh.a… <NA>
#> 11 left ar.bbr.prep lh.a… <NA>
#> 12 left atr.bbr.prep lh.a… <NA>
#> 13 left cbd.bbr.prep lh.c… <NA>
#> 14 left cbv.bbr.prep lh.c… <NA>
#> 15 left cst.bbr.prep lh.c… <NA>
#> 16 left emc.bbr.prep lh.e… <NA>
#> 17 left fat.bbr.prep lh.f… <NA>
#> 18 left fx.bbr.prep lh.f… <NA>
#> 19 left ilf.bbr.prep lh.i… <NA>
#> 20 left mlf.bbr.prep lh.m… <NA>
#> 21 left or.bbr.prep lh.o… <NA>
#> 22 left slf1.bbr.prep lh.s… <NA>
#> 23 left slf2.bbr.prep lh.s… <NA>
#> 24 left slf3.bbr.prep lh.s… <NA>
#> 25 left uf.bbr.prep lh.u… <NA>
#> 26 midline mcp.bbr.prep mcp.… <NA>
#> 27 right af.bbr.prep rh.a… <NA>
#> 28 right ar.bbr.prep rh.a… <NA>
#> 29 right atr.bbr.prep rh.a… <NA>
#> 30 right cbd.bbr.prep rh.c… <NA>
#> 31 right cbv.bbr.prep rh.c… <NA>
#> 32 right cst.bbr.prep rh.c… <NA>
#> 33 right emc.bbr.prep rh.e… <NA>
#> 34 right fat.bbr.prep rh.f… <NA>
#> 35 right fx.bbr.prep rh.f… <NA>
#> 36 right ilf.bbr.prep rh.i… <NA>
#> 37 right mlf.bbr.prep rh.m… <NA>
#> 38 right or.bbr.prep rh.o… <NA>
#> 39 right slf1.bbr.prep rh.s… <NA>
#> 40 right slf2.bbr.prep rh.s… <NA>
#> 41 right slf3.bbr.prep rh.s… <NA>
#> 42 right uf.bbr.prep rh.u… <NA>
atlas_labels(tracula)
#> [1] "acomm.bbr.prep"
#> [2] "cc.bodyc.bbr.prep"
#> [3] "cc.bodyp.bbr.prep"
#> [4] "cc.bodypf.bbr.prep"
#> [5] "cc.bodypm.bbr.prep"
#> [6] "cc.bodyt.bbr.prep"
#> [7] "cc.genu.bbr.prep"
#> [8] "cc.rostrum.bbr.prep"
#> [9] "cc.splenium.bbr.prep"
#> [10] "lh.af.bbr.prep"
#> [11] "lh.ar.bbr.prep"
#> [12] "lh.atr.bbr.prep"
#> [13] "lh.cbd.bbr.prep"
#> [14] "lh.cbv.bbr.prep"
#> [15] "lh.cst.bbr.prep"
#> [16] "lh.emc.bbr.prep"
#> [17] "lh.fat.bbr.prep"
#> [18] "lh.fx.bbr.prep"
#> [19] "lh.ilf.bbr.prep"
#> [20] "lh.mlf.bbr.prep"
#> [21] "lh.or.bbr.prep"
#> [22] "lh.slf1.bbr.prep"
#> [23] "lh.slf2.bbr.prep"
#> [24] "lh.slf3.bbr.prep"
#> [25] "lh.uf.bbr.prep"
#> [26] "mcp.bbr.prep"
#> [27] "rh.af.bbr.prep"
#> [28] "rh.ar.bbr.prep"
#> [29] "rh.atr.bbr.prep"
#> [30] "rh.cbd.bbr.prep"
#> [31] "rh.cbv.bbr.prep"
#> [32] "rh.cst.bbr.prep"
#> [33] "rh.emc.bbr.prep"
#> [34] "rh.fat.bbr.prep"
#> [35] "rh.fx.bbr.prep"
#> [36] "rh.ilf.bbr.prep"
#> [37] "rh.mlf.bbr.prep"
#> [38] "rh.or.bbr.prep"
#> [39] "rh.slf1.bbr.prep"
#> [40] "rh.slf2.bbr.prep"
#> [41] "rh.slf3.bbr.prep"
#> [42] "rh.uf.bbr.prep"
table(tracula$core$group, useNA = "ifany")
#>
#> <NA>
#> 42
plot(tracula) +
ggplot2::scale_fill_viridis_d(na.value = "grey80")
#> Scale for fill is already present.
#> Adding another scale for fill, which will
#> replace the existing scale.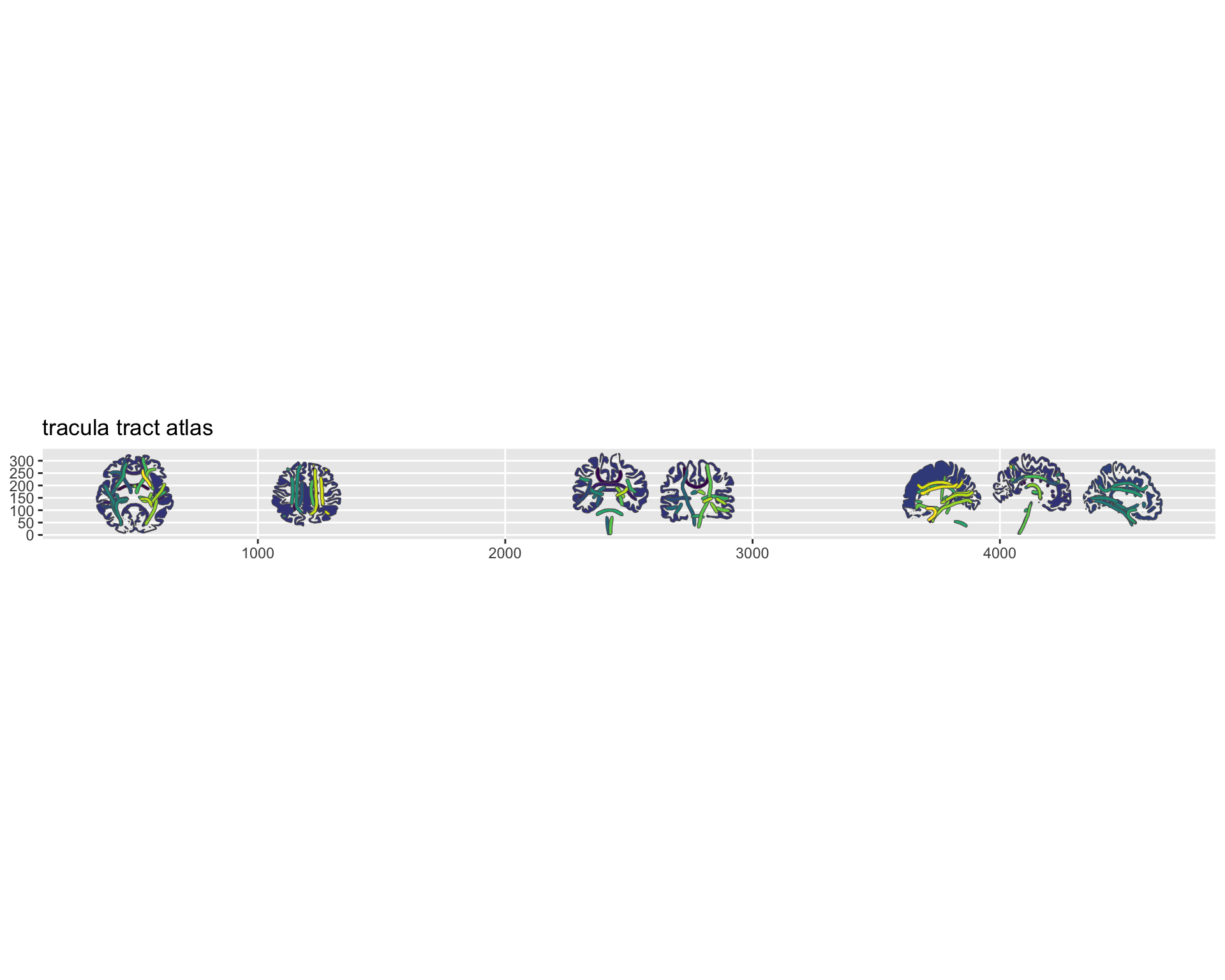
TRACULA white matter tract atlas plotted with ggseg.