Subcortical atlases represent the structures beneath the cortical surface: thalamus, hippocampus, amygdala, caudate, putamen, and others. They come from volumetric segmentations like FreeSurfer’s aseg.mgz, where each voxel is assigned a label.
The pipeline tessellates labelled voxel regions into 3D meshes and (optionally) creates 2D projection views that collapse a slab of the volume onto a single plane.
This tutorial recreates the aseg atlas — the same pipeline behind data-raw/make_aseg_atlas.R in ggseg.formats.
What you need
- FreeSurfer installed with the
fsaverage5subject - ImageMagick for 2D geometry extraction
- Chrome/Chromium for 3D screenshots
Creating the atlas
create_subcortical_from_volume() takes a segmentation volume and a colour lookup table. The pipeline tessellates each labelled region into a 3D mesh, then creates 2D projection views:
aseg_raw <- create_subcortical_from_volume(
input_volume = aseg_volume,
input_lut = color_lut,
atlas_name = "aseg"
)
#> Warning: Atlas has 11943 vertices (threshold:
#> 10000)
#> ℹ Large atlases may be slow to plot and
#> increase package size
#> ℹ Re-run with higher `tolerance` to
#> reduce vertices
aseg_raw
#>
#> ── aseg ggseg atlas ──────────────────────
#> Type: subcortical
#> Regions: 27
#> Hemispheres: left, NA, right
#> Views: axial_1, axial_2, axial_3,
#> axial_4, axial_5, axial_6, axial_7,
#> coronal_1, coronal_2, coronal_3,
#> coronal_4, coronal_5, sagittal
#> Palette: ✔
#> Rendering: ✔ ggseg
#> ✔ ggseg3d (meshes)
#> ──────────────────────────────────────────
#> # A tibble: 43 × 3
#> hemi region label
#> <chr> <chr> <chr>
#> 1 left cerebral white matter Left-Cer…
#> 2 left cerebral cortex Left-Cer…
#> 3 left lateral ventricle Left-Lat…
#> 4 left inf lat vent Left-Inf…
#> 5 left cerebellum white matter Left-Cer…
#> 6 left cerebellum cortex Left-Cer…
#> 7 left thalamus Left-Tha…
#> 8 left caudate Left-Cau…
#> 9 left putamen Left-Put…
#> 10 left pallidum Left-Pal…
#> 11 <NA> 3rd ventricle 3rd-Vent…
#> 12 <NA> 4th ventricle 4th-Vent…
#> 13 <NA> brain stem Brain-St…
#> 14 left hippocampus Left-Hip…
#> 15 left amygdala Left-Amy…
#> 16 <NA> csf CSF
#> 17 left accumbens area Left-Acc…
#> 18 left ventraldc Left-Ven…
#> 19 left vessel Left-ves…
#> 20 left choroid plexus Left-cho…
#> 21 right cerebral white matter Right-Ce…
#> 22 right cerebral cortex Right-Ce…
#> 23 right lateral ventricle Right-La…
#> 24 right inf lat vent Right-In…
#> 25 right cerebellum white matter Right-Ce…
#> 26 right cerebellum cortex Right-Ce…
#> 27 right thalamus Right-Th…
#> 28 right caudate Right-Ca…
#> 29 right putamen Right-Pu…
#> 30 right pallidum Right-Pa…
#> 31 right hippocampus Right-Hi…
#> 32 right amygdala Right-Am…
#> 33 right accumbens area Right-Ac…
#> 34 right ventraldc Right-Ve…
#> 35 right vessel Right-ve…
#> 36 right choroid plexus Right-ch…
#> 37 <NA> wm hypointensities WM-hypoi…
#> 38 <NA> optic chiasm Optic-Ch…
#> 39 <NA> cc posterior CC_Poste…
#> 40 <NA> cc mid posterior CC_Mid_P…
#> 41 <NA> cc central CC_Centr…
#> 42 <NA> cc mid anterior CC_Mid_A…
#> 43 <NA> cc anterior CC_Anter…The default pipeline creates six projection views focused on the subcortical range: axial inferior, axial superior, coronal posterior, coronal anterior, sagittal left, and sagittal right. Each view collapses a slab of the volume onto a single plane, giving you spatial context without the complexity of individual slices.
Removing unwanted regions
The aseg segmentation contains everything — cortex, white matter, ventricles, CSF. For a subcortical atlas, most of these are noise. Remove them:
aseg_raw <- aseg_raw |>
atlas_region_remove("White-Matter", match_on = "label") |>
atlas_region_remove("WM-hypointensities", match_on = "label") |>
atlas_region_remove("-Ventricle", match_on = "label") |>
atlas_region_remove("-Vent$", match_on = "label") |>
atlas_region_remove("CSF", match_on = "label") |>
atlas_region_remove("Cerebral-Cortex", match_on = "label")Patterns are regular expressions, so -Vent$ matches “3rd-Vent” and “4th-Vent” without catching “Ventral-DC.”
Setting context regions
The cortex works well as a background outline — it shows where subcortical structures sit relative to the brain surface without competing for colour:
aseg_raw <- aseg_raw |>
atlas_region_contextual("Cortex", match_on = "label")Selecting views
Not all projection views are equally useful. Keep the ones that show your structures best:
aseg_raw <- aseg_raw |>
atlas_view_keep("axial_3|axial_5|coronal_2|coronal_3|coronal_4|sagittal")Cleaning up the layout
Gather views into a compact arrangement:
aseg_raw <- aseg_raw |>
atlas_view_gather()Adding metadata
The raw labels are technical identifiers like “Left-Thalamus-Proper.” Join human-readable names and structural groupings:
normalize_region <- function(x) {
ifelse(
is.na(x),
NA_character_,
x |>
tolower() |>
gsub("-", " ", x = _) |>
gsub("_", " ", x = _) |>
gsub("left |right ", "", x = _) |>
trimws()
)
}
aseg_metadata <- data.frame(
region = c(
"Thalamus-Proper", "Caudate", "Putamen", "Pallidum",
"Hippocampus", "Amygdala", "Accumbens-area", "VentralDC",
"Brain-Stem"
),
label_pretty = c(
"thalamus", "caudate", "putamen", "pallidum",
"hippocampus", "amygdala", "nucleus accumbens", "ventral DC",
"brain stem"
),
structure = c(
"diencephalon", "basal ganglia", "basal ganglia", "basal ganglia",
"limbic", "limbic", "basal ganglia", "diencephalon",
"brainstem"
)
)
core_with_meta <- aseg_raw$core |>
mutate(region_key = normalize_region(region)) |>
left_join(
aseg_metadata |>
mutate(region_key = normalize_region(region)) |>
select(region_key, label_pretty, structure),
by = "region_key"
) |>
mutate(region = coalesce(label_pretty, region)) |>
select(hemi, region, label, structure)Rebuilding and saving
Construct the final atlas from modified components:
aseg <- ggseg_atlas(
atlas = aseg_raw$atlas,
type = aseg_raw$type,
palette = aseg_raw$palette,
core = core_with_meta,
data = aseg_raw$data
)
aseg
#>
#> ── aseg ggseg atlas ──────────────────────
#> Type: subcortical
#> Regions: 17
#> Hemispheres: left, NA, right
#> Views: axial_3, axial_5, coronal_2,
#> coronal_3, coronal_4, sagittal
#> Palette: ✔
#> Rendering: ✔ ggseg
#> ✔ ggseg3d (meshes)
#> ──────────────────────────────────────────
#> # A tibble: 27 × 4
#> hemi region label structure
#> <chr> <chr> <chr> <chr>
#> 1 left thalamus Left… <NA>
#> 2 left caudate Left… basal ga…
#> 3 left putamen Left… basal ga…
#> 4 left pallidum Left… basal ga…
#> 5 <NA> brain stem Brai… brainstem
#> 6 left hippocampus Left… limbic
#> 7 left amygdala Left… limbic
#> 8 left nucleus accumbens Left… basal ga…
#> 9 left ventral DC Left… dienceph…
#> 10 left vessel Left… <NA>
#> 11 left choroid plexus Left… <NA>
#> 12 right thalamus Righ… <NA>
#> 13 right caudate Righ… basal ga…
#> 14 right putamen Righ… basal ga…
#> 15 right pallidum Righ… basal ga…
#> 16 right hippocampus Righ… limbic
#> 17 right amygdala Righ… limbic
#> 18 right nucleus accumbens Righ… basal ga…
#> 19 right ventral DC Righ… dienceph…
#> 20 right vessel Righ… <NA>
#> 21 right choroid plexus Righ… <NA>
#> 22 <NA> optic chiasm Opti… <NA>
#> 23 <NA> cc posterior CC_P… <NA>
#> 24 <NA> cc mid posterior CC_M… <NA>
#> 25 <NA> cc central CC_C… <NA>
#> 26 <NA> cc mid anterior CC_M… <NA>
#> 27 <NA> cc anterior CC_A… <NA>
atlas_labels(aseg)
#> [1] "Brain-Stem"
#> [2] "CC_Anterior"
#> [3] "CC_Central"
#> [4] "CC_Mid_Anterior"
#> [5] "CC_Mid_Posterior"
#> [6] "CC_Posterior"
#> [7] "Left-Accumbens-area"
#> [8] "Left-Amygdala"
#> [9] "Left-Caudate"
#> [10] "Left-choroid-plexus"
#> [11] "Left-Hippocampus"
#> [12] "Left-Pallidum"
#> [13] "Left-Putamen"
#> [14] "Left-Thalamus"
#> [15] "Left-VentralDC"
#> [16] "Left-vessel"
#> [17] "Optic-Chiasm"
#> [18] "Right-Accumbens-area"
#> [19] "Right-Amygdala"
#> [20] "Right-Caudate"
#> [21] "Right-choroid-plexus"
#> [22] "Right-Hippocampus"
#> [23] "Right-Pallidum"
#> [24] "Right-Putamen"
#> [25] "Right-Thalamus"
#> [26] "Right-VentralDC"
#> [27] "Right-vessel"
table(aseg$core$structure)
#>
#> basal ganglia brainstem diencephalon
#> 8 1 2
#> limbic
#> 4
plot(aseg)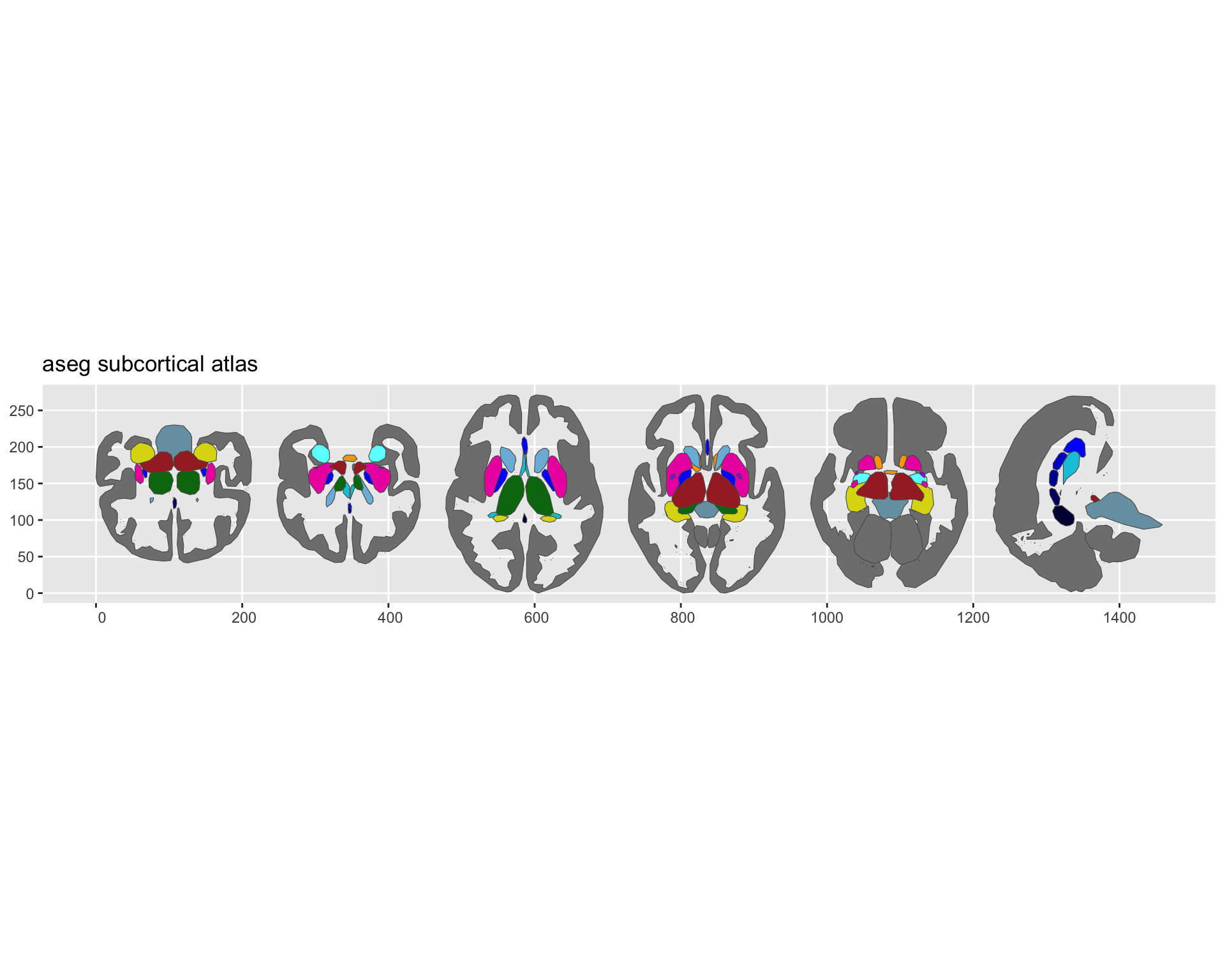
Subcortical aseg atlas plotted with ggseg.