geom_brain() handles data joining and positioning
automatically, which covers most use cases. But because it owns the data
pipeline, you can’t easily layer other sf geoms on top – things like
geom_sf_label() or geom_sf_text() need direct
access to the sf data.
This vignette shows how to work with brain atlases as plain sf objects, giving you full control at the cost of a few extra lines.
library(ggseg)
library(ggplot2)
library(dplyr)
#>
#> Attaching package: 'dplyr'
#> The following objects are masked from 'package:stats':
#>
#> filter, lag
#> The following objects are masked from 'package:base':
#>
#> intersect, setdiff, setequal, unionWhen you’d want this
Use the sf workflow when you need to:
- Add region labels with
geom_sf_label()orgeom_sf_text() - Use
ggrepel::geom_label_repel()for non-overlapping labels - Layer other sf geoms on the brain
- Have full control over the data before it reaches ggplot
For everything else, geom_brain() is simpler.
Getting atlas data as sf
A brain atlas stores its geometry in the data slot.
Convert to a plain data frame to see all the columns:
dk()$data
#>
#> ── ggseg_data_cortical ──
#>
#> 2D (ggseg): 72 labels, views: inferior, lateral, medial, superior
#> 3D (ggseg3d): vertex indices
#> # A tibble: 70 × 2
#> label vertices
#> <chr> <list>
#> 1 lh_bankssts <int [126]>
#> 2 lh_caudalanteriorcingulate <int [67]>
#> 3 lh_caudalmiddlefrontal <int [232]>
#> 4 lh_corpuscallosum <int [198]>
#> 5 lh_cuneus <int [102]>
#> 6 lh_entorhinal <int [48]>
#> 7 lh_fusiform <int [308]>
#> 8 lh_inferiorparietal <int [484]>
#> 9 lh_inferiortemporal <int [271]>
#> 10 lh_isthmuscingulate <int [123]>
#> # ℹ 60 more rowsThe geometry column holds the polygons. This is a
standard sf object, so any sf-compatible tool works with it.
Joining your data
Use brain_join() to merge your data with the atlas. It
preserves the sf geometry and handles column detection:
some_data <- tibble(
region = c(
"transverse temporal",
"insula",
"precentral",
"superior parietal"
),
p = sample(seq(0, 0.5, 0.001), 4)
)
some_data |>
brain_join(dk())
#> Merging atlas and data by region.
#> Simple feature collection with 191 features and 9 fields
#> Geometry type: MULTIPOLYGON
#> Dimension: XY
#> Bounding box: xmin: 84.2049 ymin: 0 xmax: 5359.689 ymax: 429.9372
#> CRS: NA
#> First 10 features:
#> label view hemi region
#> 1 lh_unknown medial left <NA>
#> 2 lh_unknown lateral left <NA>
#> 3 lh_unknown inferior left <NA>
#> 4 rh_unknown lateral right <NA>
#> 5 rh_unknown inferior right <NA>
#> 6 rh_unknown medial right <NA>
#> 7 lh_bankssts inferior left banks of superior temporal sulcus
#> 8 lh_bankssts lateral left banks of superior temporal sulcus
#> 9 lh_bankssts superior left banks of superior temporal sulcus
#> 10 lh_caudalanteriorcingulate medial left caudal anterior cingulate
#> lobe atlas type colour p geometry
#> 1 <NA> dk cortical <NA> NA MULTIPOLYGON (((1782.84 18....
#> 2 <NA> dk cortical <NA> NA MULTIPOLYGON (((926.5936 60...
#> 3 <NA> dk cortical <NA> NA MULTIPOLYGON (((367.1256 13...
#> 4 <NA> dk cortical <NA> NA MULTIPOLYGON (((3849.766 60...
#> 5 <NA> dk cortical <NA> NA MULTIPOLYGON (((3190.519 5....
#> 6 <NA> dk cortical <NA> NA MULTIPOLYGON (((4318.844 20...
#> 7 temporal dk cortical #196428 NA MULTIPOLYGON (((534.4782 21...
#> 8 temporal dk cortical #196428 NA MULTIPOLYGON (((1121.478 12...
#> 9 temporal dk cortical #196428 NA MULTIPOLYGON (((2448.464 20...
#> 10 cingulate dk cortical #7D64A0 NA MULTIPOLYGON (((1921.971 20...The result is a standard sf object you can pass to
geom_sf().
Plotting with geom_sf
some_data |>
brain_join(dk()) |>
ggplot() +
geom_sf(aes(fill = p))
#> Merging atlas and data by region.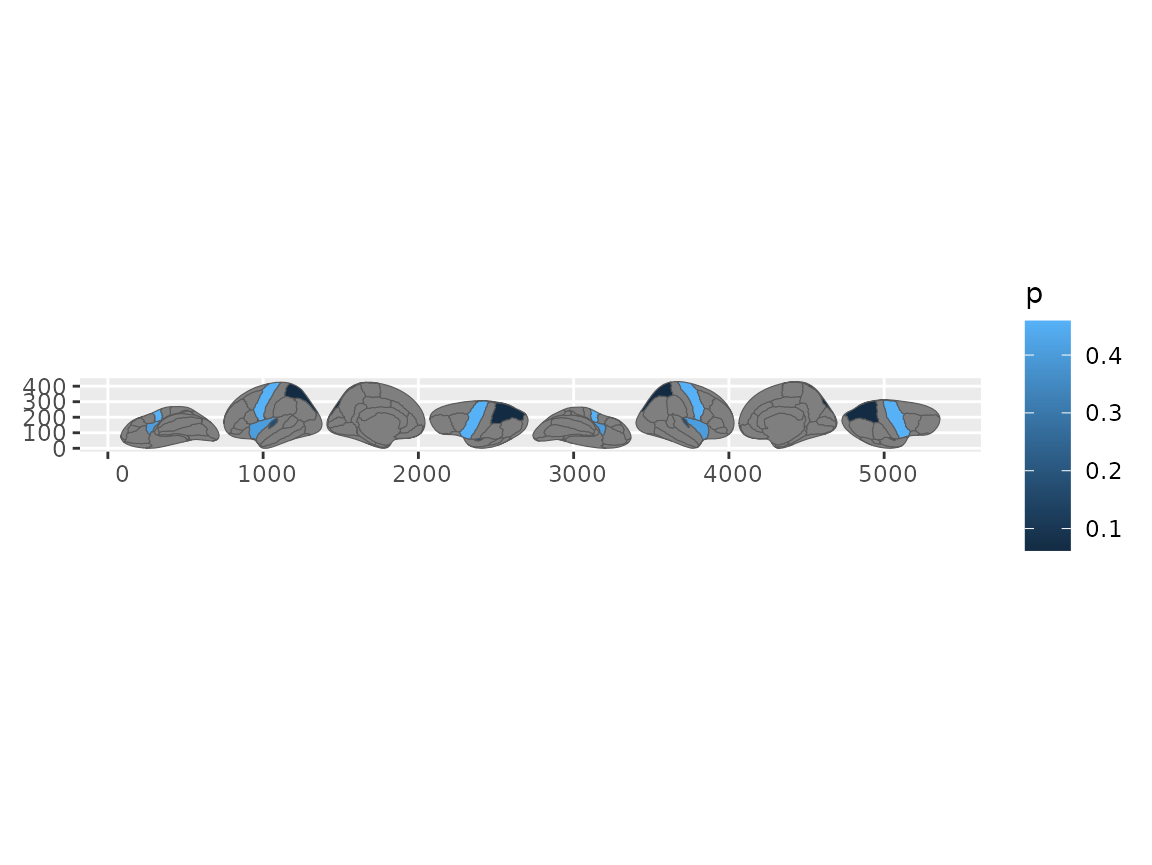
Brain plot using geom_sf after brain_join.
Repositioning views
With geom_brain(), you’d use
position_brain(). In the sf workflow, use
reposition_brain() instead – it transforms the geometry
directly:
some_data |>
brain_join(dk()) |>
reposition_brain(hemi ~ view) |>
ggplot() +
geom_sf(aes(fill = p))
#> Merging atlas and data by region.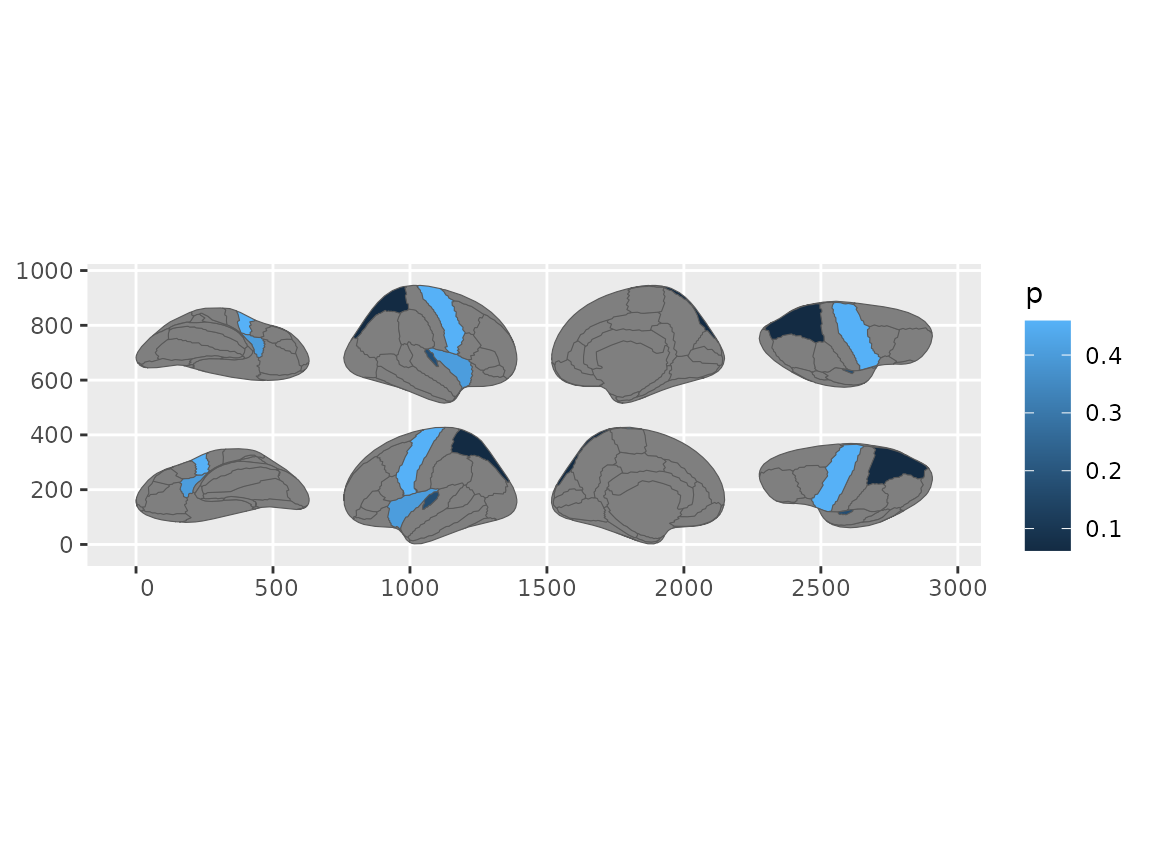
Repositioned brain views using reposition_brain() with geom_sf.
Same formula syntax, same results.
Adding labels
This is the main reason to use the sf workflow. Once you have repositioned sf data, you can layer any sf geom:
some_data |>
brain_join(dk()) |>
reposition_brain(hemi ~ view) |>
ggplot(aes(fill = p)) +
geom_sf(show.legend = FALSE) +
geom_sf_label(
aes(label = ifelse(!is.na(p), region, NA)),
alpha = 0.8,
show.legend = FALSE
)
#> Merging atlas and data by region.
#> Warning: Removed 168 rows containing missing values or values outside the scale range
#> (`geom_label()`).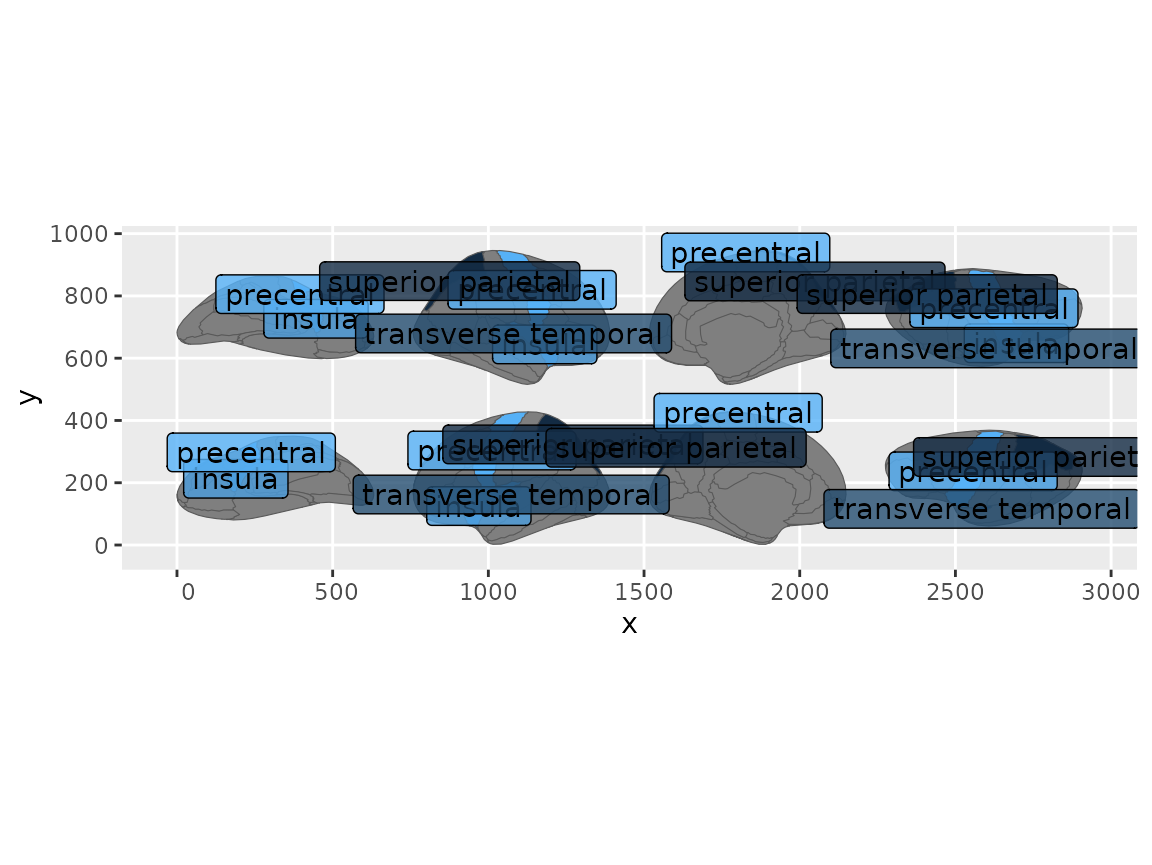
Brain regions with text labels overlaid using geom_sf_label.
For crowded plots, ggrepel::geom_label_repel() avoids
overlapping labels.
Faceting with grouped data
In the sf workflow, faceting requires group_by() before
brain_join(). The grouping tells the join to replicate the
atlas for each group:
some_data <- tibble(
region = rep(
c(
"transverse temporal",
"insula",
"precentral",
"superior parietal"
),
2
),
p = sample(seq(0, 0.5, 0.001), 8),
group = c(rep("A", 4), rep("B", 4))
)
some_data |>
group_by(group) |>
brain_join(dk()) |>
reposition_brain(hemi ~ view) |>
ggplot(aes(fill = p)) +
geom_sf(show.legend = FALSE) +
facet_wrap(~group)
#> Merging atlas and data by region.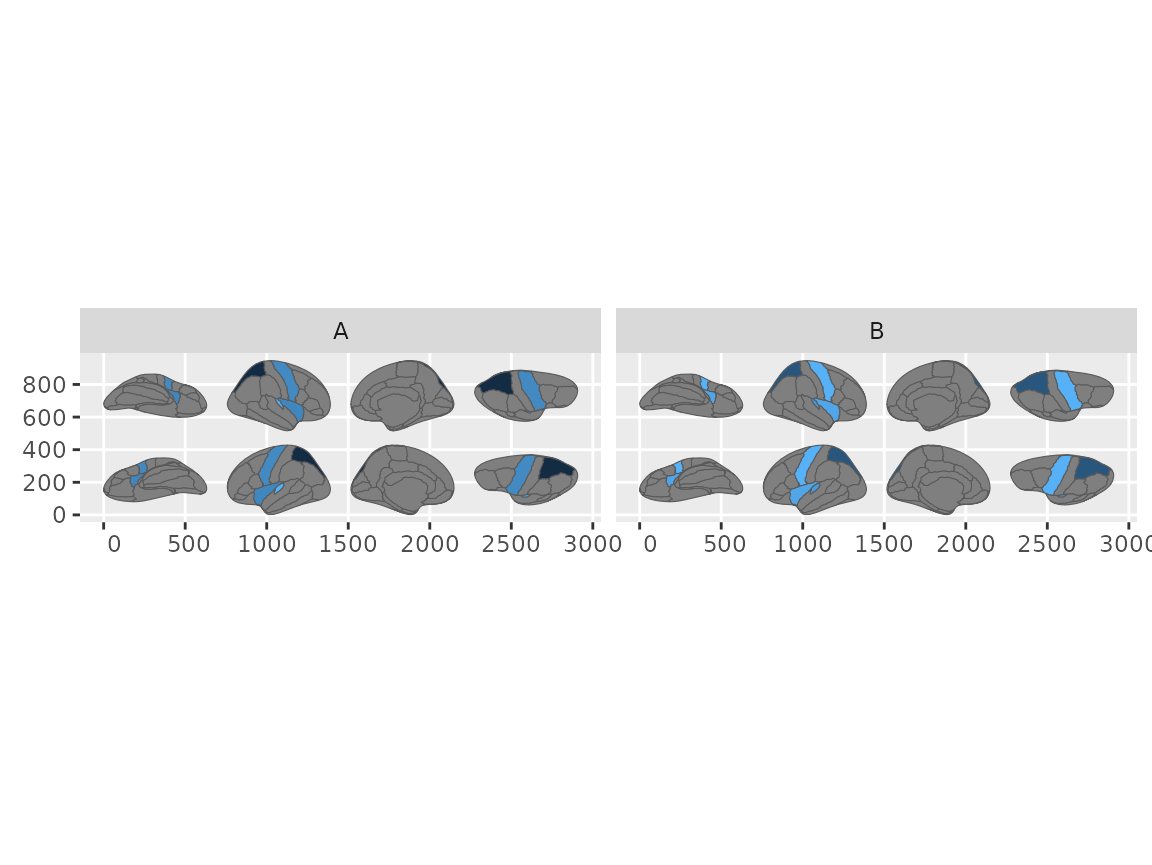
Faceted brain plots using the geom_sf workflow with grouped data.
This step is only needed in the sf workflow.
geom_brain() handles atlas replication automatically (see
vignette("external-data")).
