If you work with neuroimaging data, you’ve probably spent time wrestling region-level results into brain plots. ggseg handles that plumbing for you. It stores brain atlas geometries as simple features and plots them as ggplot2 layers, so you get brain figures with the same code you’d use for any other ggplot.
What’s in a brain atlas
A brain_atlas object bundles four things: a name, a type
(cortical, subcortical, or tract), region geometry stored as simple
features, and an optional colour palette.
dk
#> function ()
#> .dk_atlas
#> <bytecode: 0x55ea66fed3f0>
#> <environment: namespace:ggseg.formats>The dk atlas (Desikan-Killiany) ships with the package.
Call plot() for a quick look:
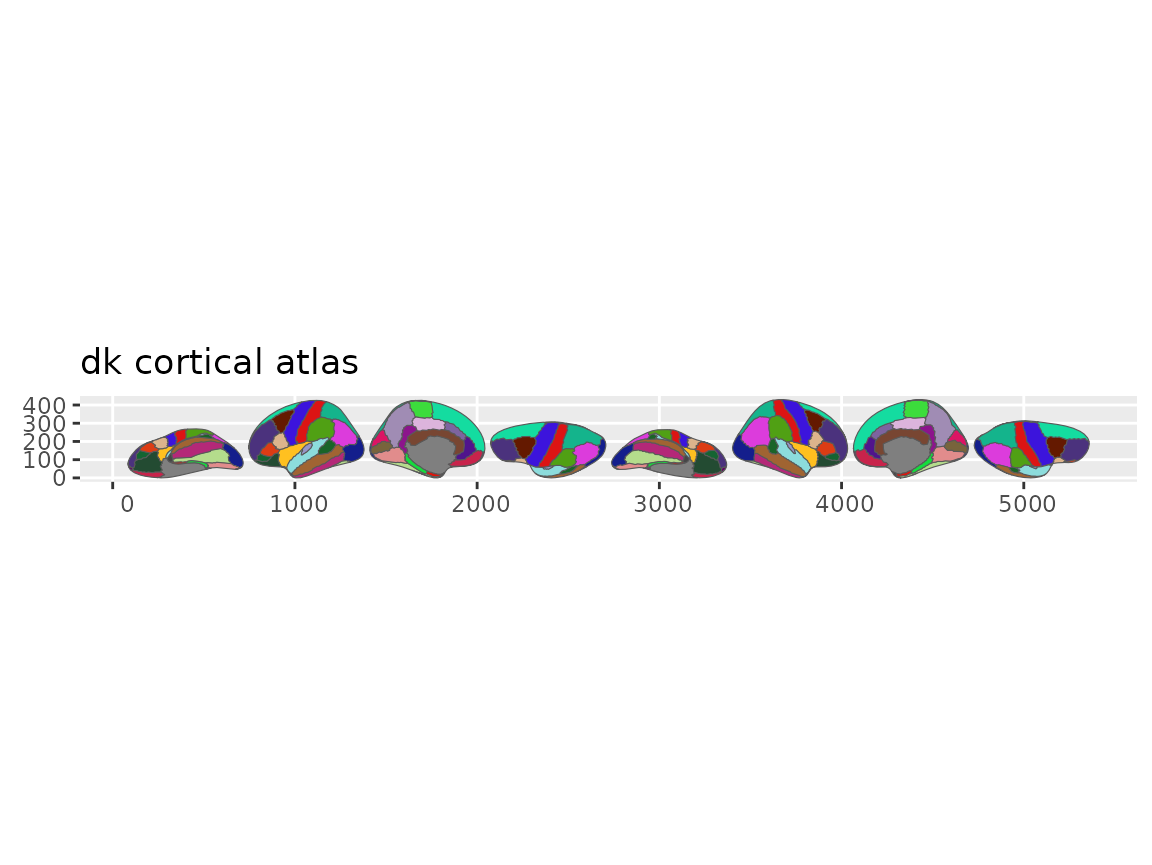
Quick overview of the Desikan-Killiany cortical atlas.
Two helpers pull out the names you’ll need for matching data:
ggseg.formats::atlas_regions(dk())
#> [1] "banks of superior temporal sulcus" "caudal anterior cingulate"
#> [3] "caudal middle frontal" "corpus callosum"
#> [5] "cuneus" "entorhinal"
#> [7] "frontal pole" "fusiform"
#> [9] "inferior parietal" "inferior temporal"
#> [11] "insula" "isthmus cingulate"
#> [13] "lateral occipital" "lateral orbitofrontal"
#> [15] "lingual" "medial orbitofrontal"
#> [17] "middle temporal" "paracentral"
#> [19] "parahippocampal" "pars opercularis"
#> [21] "pars orbitalis" "pars triangularis"
#> [23] "pericalcarine" "postcentral"
#> [25] "posterior cingulate" "precentral"
#> [27] "precuneus" "rostral anterior cingulate"
#> [29] "rostral middle frontal" "superior frontal"
#> [31] "superior parietal" "superior temporal"
#> [33] "supramarginal" "temporal pole"
#> [35] "transverse temporal"
ggseg.formats::atlas_labels(dk())
#> [1] "lh_bankssts" "lh_caudalanteriorcingulate"
#> [3] "lh_caudalmiddlefrontal" "lh_corpuscallosum"
#> [5] "lh_cuneus" "lh_entorhinal"
#> [7] "lh_frontalpole" "lh_fusiform"
#> [9] "lh_inferiorparietal" "lh_inferiortemporal"
#> [11] "lh_insula" "lh_isthmuscingulate"
#> [13] "lh_lateraloccipital" "lh_lateralorbitofrontal"
#> [15] "lh_lingual" "lh_medialorbitofrontal"
#> [17] "lh_middletemporal" "lh_paracentral"
#> [19] "lh_parahippocampal" "lh_parsopercularis"
#> [21] "lh_parsorbitalis" "lh_parstriangularis"
#> [23] "lh_pericalcarine" "lh_postcentral"
#> [25] "lh_posteriorcingulate" "lh_precentral"
#> [27] "lh_precuneus" "lh_rostralanteriorcingulate"
#> [29] "lh_rostralmiddlefrontal" "lh_superiorfrontal"
#> [31] "lh_superiorparietal" "lh_superiortemporal"
#> [33] "lh_supramarginal" "lh_temporalpole"
#> [35] "lh_transversetemporal" "rh_bankssts"
#> [37] "rh_caudalanteriorcingulate" "rh_caudalmiddlefrontal"
#> [39] "rh_corpuscallosum" "rh_cuneus"
#> [41] "rh_entorhinal" "rh_frontalpole"
#> [43] "rh_fusiform" "rh_inferiorparietal"
#> [45] "rh_inferiortemporal" "rh_insula"
#> [47] "rh_isthmuscingulate" "rh_lateraloccipital"
#> [49] "rh_lateralorbitofrontal" "rh_lingual"
#> [51] "rh_medialorbitofrontal" "rh_middletemporal"
#> [53] "rh_paracentral" "rh_parahippocampal"
#> [55] "rh_parsopercularis" "rh_parsorbitalis"
#> [57] "rh_parstriangularis" "rh_pericalcarine"
#> [59] "rh_postcentral" "rh_posteriorcingulate"
#> [61] "rh_precentral" "rh_precuneus"
#> [63] "rh_rostralanteriorcingulate" "rh_rostralmiddlefrontal"
#> [65] "rh_superiorfrontal" "rh_superiorparietal"
#> [67] "rh_superiortemporal" "rh_supramarginal"
#> [69] "rh_temporalpole" "rh_transversetemporal"Your first brain plot
geom_brain() works like any ggplot2 geom. Pass it an
atlas and it draws the regions:
ggplot() +
geom_brain(atlas = dk())
Default brain plot using geom_brain() with the dk atlas.
Arranging views
Brain atlases have multiple views (lateral, medial) and hemispheres.
position_brain() controls how they’re laid out, using
formula syntax borrowed from facet_grid():
ggplot() +
geom_brain(atlas = dk(), position = position_brain(hemi ~ view))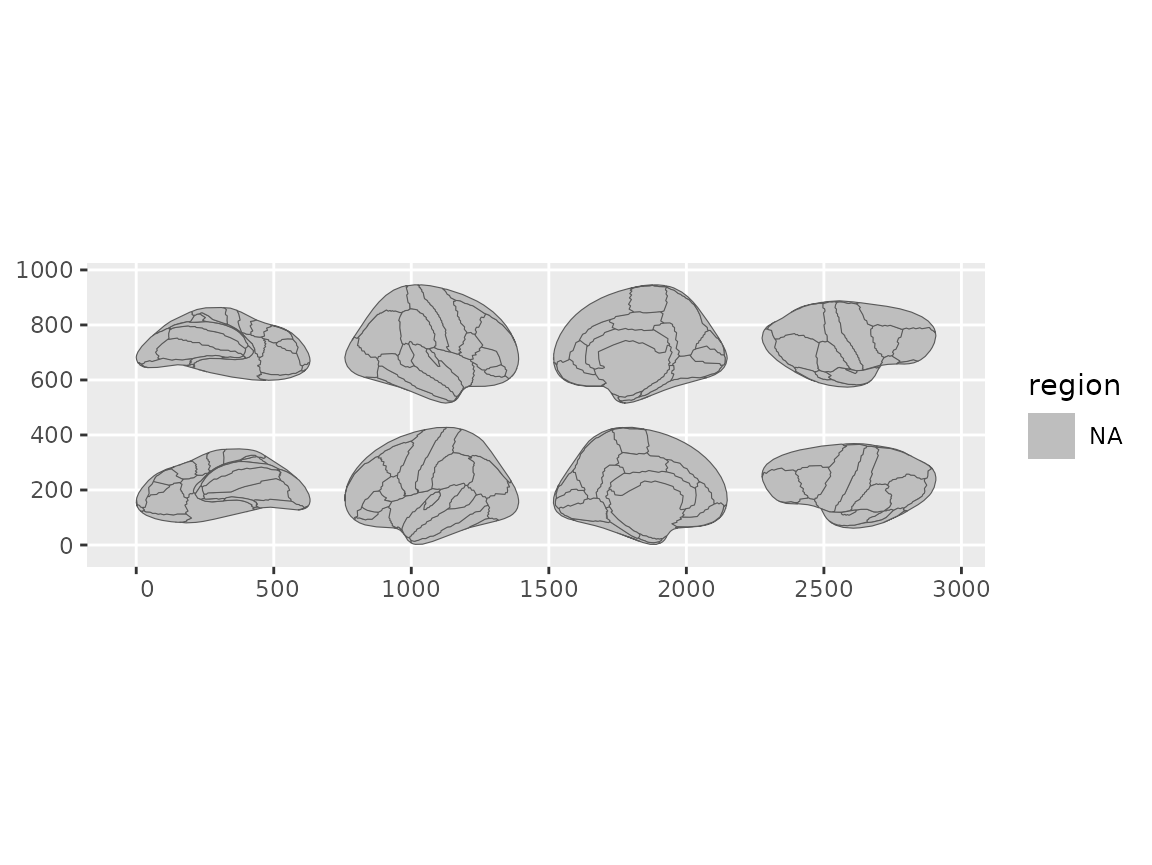
Brain views arranged using formula syntax in position_brain().
Or reorder views with a character vector:
ggplot() +
geom_brain(
atlas = dk(),
position = position_brain(c(
"right lateral",
"right medial",
"left lateral",
"left medial"
))
)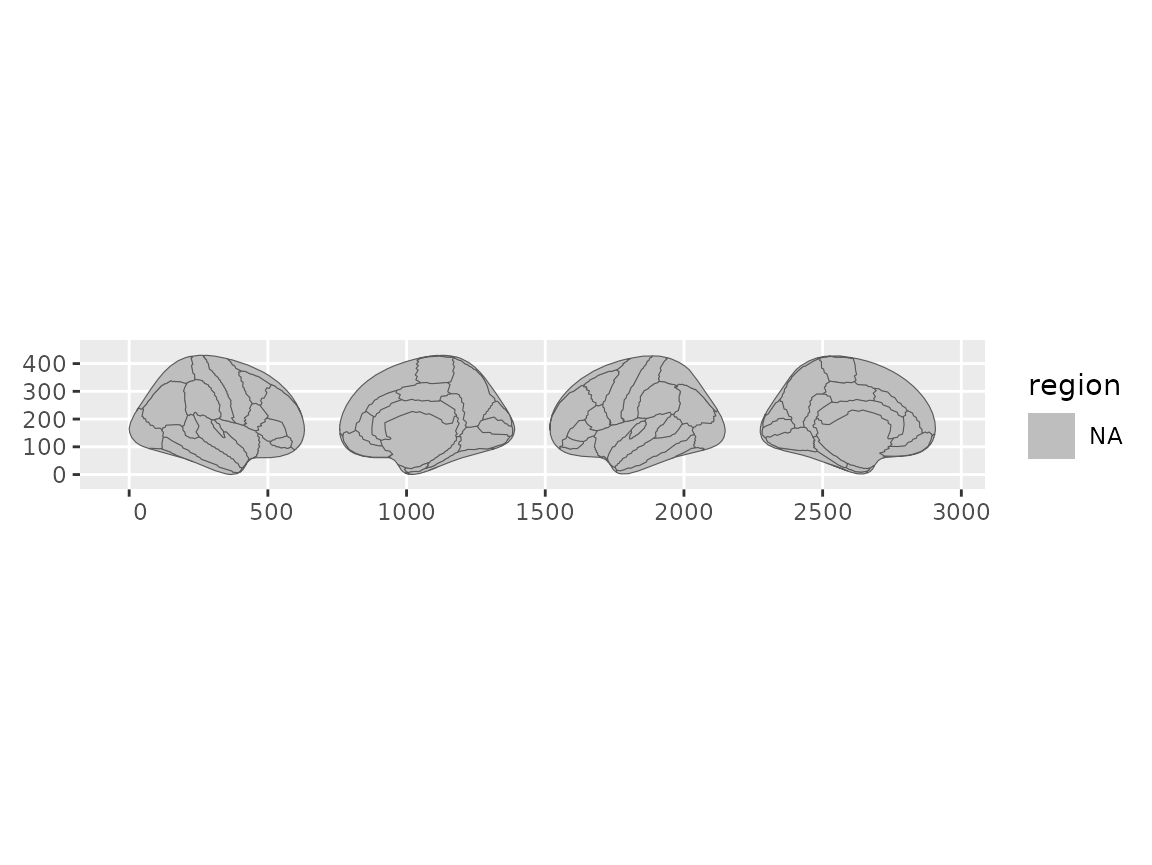
Brain views arranged using a character vector to specify order.
See vignette("positioning-views") for the full set of
layout options, including grid layouts for subcortical and tract
atlases.
Showing only certain hemispheres or views
ggplot() +
geom_brain(atlas = dk(), view = "lateral")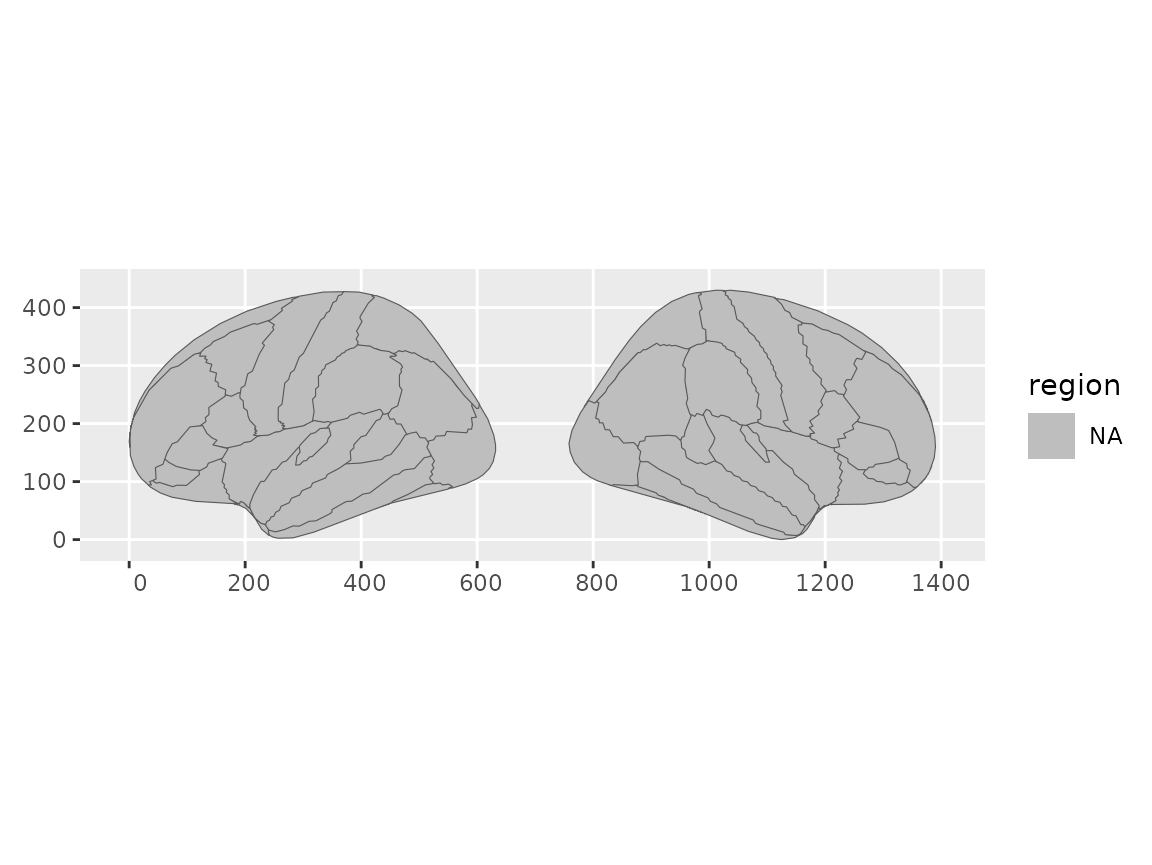
Brain plot showing only lateral views.
ggplot() +
geom_brain(atlas = dk(), hemi = "left")
Brain plot showing only the left hemisphere.
For subcortical atlases, views are slice identifiers. Check what’s
available with ggseg.formats::atlas_views():
ggseg.formats::atlas_views(aseg())
#> [1] "axial_3" "axial_4" "axial_5" "axial_6" "coronal_1" "coronal_2"
#> [7] "sagittal"Plotting your own data
Create a data frame with at least one column that matches the atlas
(typically region or label).
geom_brain() joins your data to the atlas
automatically:
library(dplyr)
#>
#> Attaching package: 'dplyr'
#> The following objects are masked from 'package:stats':
#>
#> filter, lag
#> The following objects are masked from 'package:base':
#>
#> intersect, setdiff, setequal, union
some_data <- tibble(
region = c(
"transverse temporal",
"insula",
"precentral",
"superior parietal"
),
p = sample(seq(0, 0.5, 0.001), 4)
)
ggplot(some_data) +
geom_brain(
atlas = dk(),
position = position_brain(hemi ~ view),
aes(fill = p)
) +
scale_fill_viridis_c(option = "cividis", direction = -1) +
theme_void()
#> Merging atlas and data by region.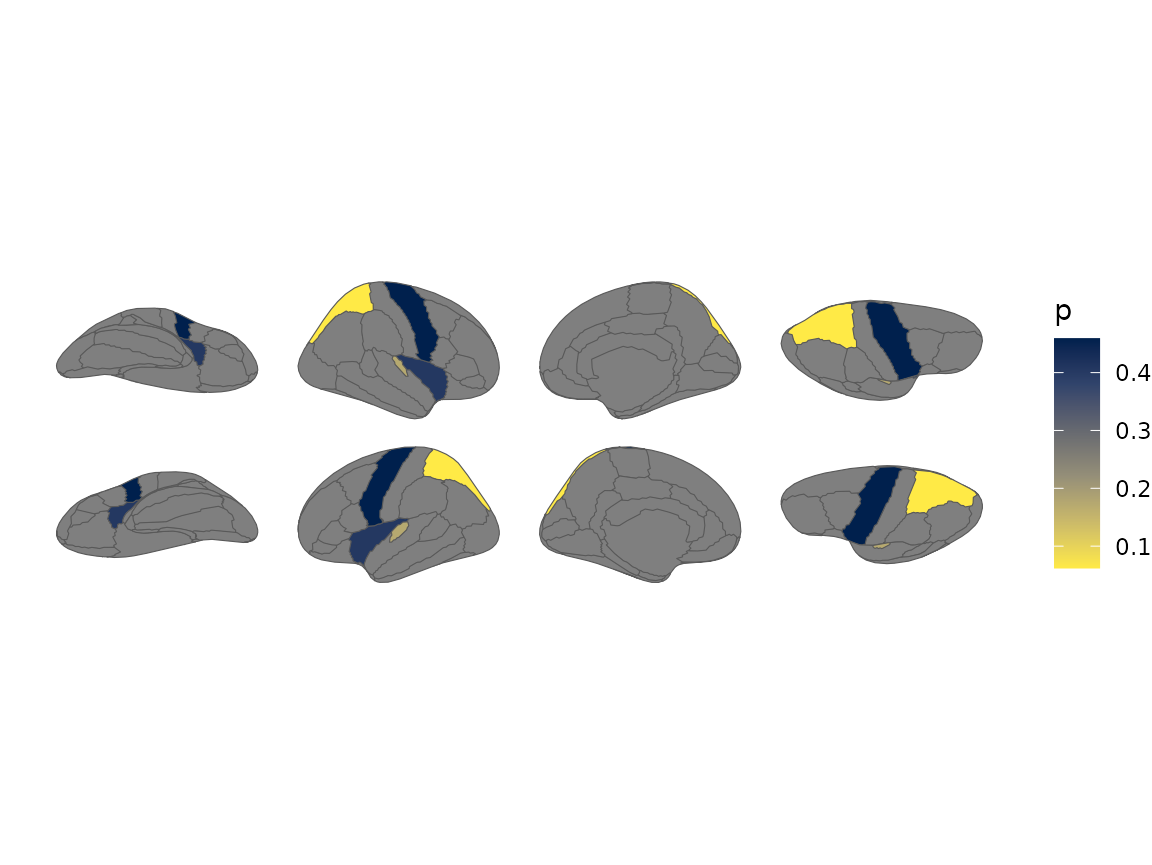
Brain plot coloured by external p-values using a viridis scale.
See vignette("external-data") for more on data
preparation and matching.
Faceting
ggplot2 faceting works as you’d expect. geom_brain()
detects faceting variables and replicates the full atlas in each
panel:
some_data <- tibble(
region = rep(
c(
"transverse temporal",
"insula",
"precentral",
"superior parietal"
),
2
),
p = sample(seq(0, 0.5, 0.001), 8),
group = c(rep("A", 4), rep("B", 4))
)
ggplot(some_data) +
geom_brain(
atlas = dk(),
position = position_brain(hemi ~ view),
aes(fill = p)
) +
facet_wrap(~group)
#> Merging atlas and data by region.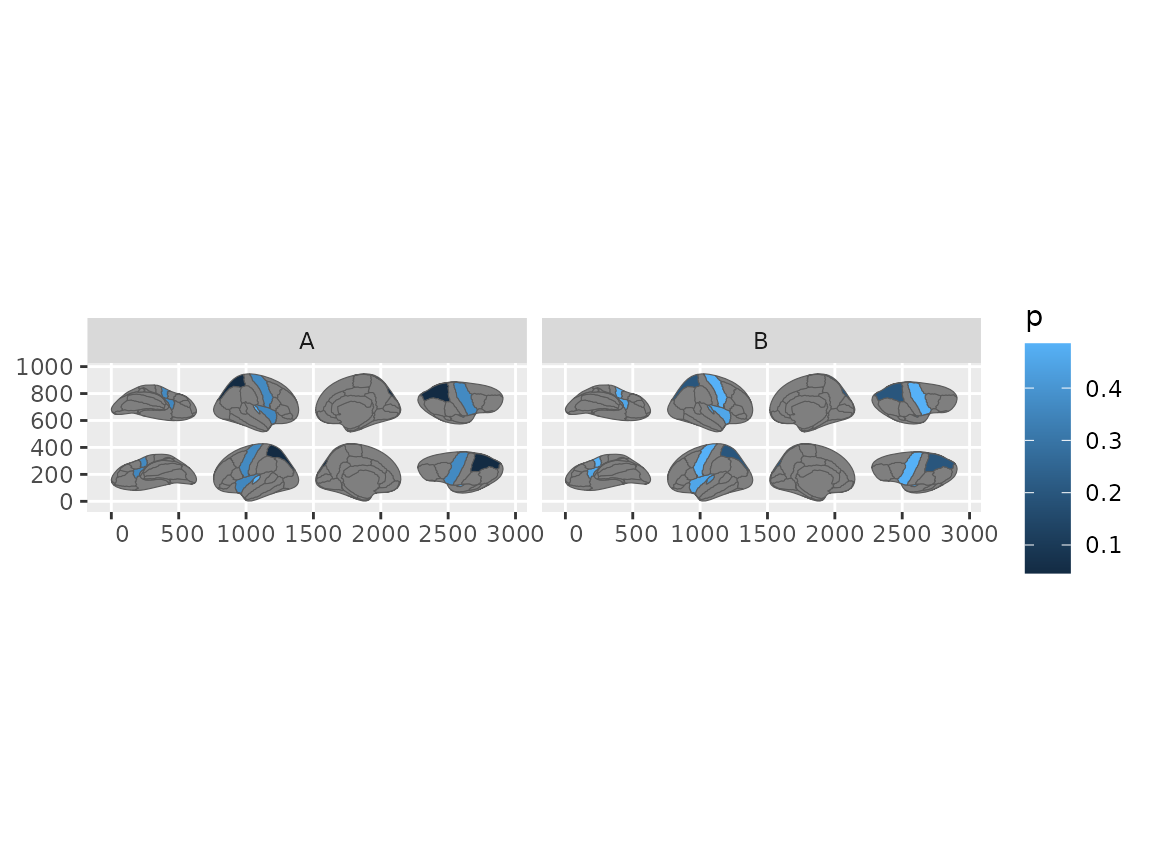
Brain plots faceted by group, each panel showing the full atlas.
No group_by() needed – the geom handles it.
Highlighting specific regions
To colour a few regions with their atlas colours, use two columns:
one for the join (region) and one for the fill
aesthetic:
data <- data.frame(
region = ggseg.formats::atlas_regions(dk())[1:3],
reg_col = ggseg.formats::atlas_regions(dk())[1:3]
)
ggplot(data) +
geom_brain(atlas = dk(), aes(fill = reg_col)) +
scale_fill_brain_manual(dk()$palette[data$region])
#> Merging atlas and data by region.
Highlighting selected regions using the atlas colour palette.
Scales and themes
Atlas colour palettes are applied automatically by
geom_brain(). For custom palettes, use
scale_fill_brain_manual().
Built-in themes strip axes and grids for cleaner figures:
ggplot() +
geom_brain(atlas = dk()) +
theme_brain()Finding more atlases
ggseg ships with dk (cortical), aseg
(subcortical), and tracula (white matter tracts). Many more
are available through the ggsegverse r-universe:
install.packages("ggsegYeo2011", repos = "https://ggsegverse.r-universe.dev")