A brain atlas typically has several views – lateral and medial for
cortical atlases, or axial, coronal, and sagittal slices for subcortical
and tract atlases. position_brain() controls how those
views are arranged in the final plot.
The function works differently depending on the atlas type, so this vignette covers cortical and subcortical/tract atlases separately.
Cortical atlases
Cortical atlases like dk have two layout dimensions:
hemi (left or right) and view
(lateral, medial, etc.). The formula syntax mirrors
facet_grid() – left side is rows, right side is
columns:
ggplot() +
geom_brain(
atlas = dk(),
position = position_brain(hemi ~ view),
show.legend = FALSE
) +
theme_void()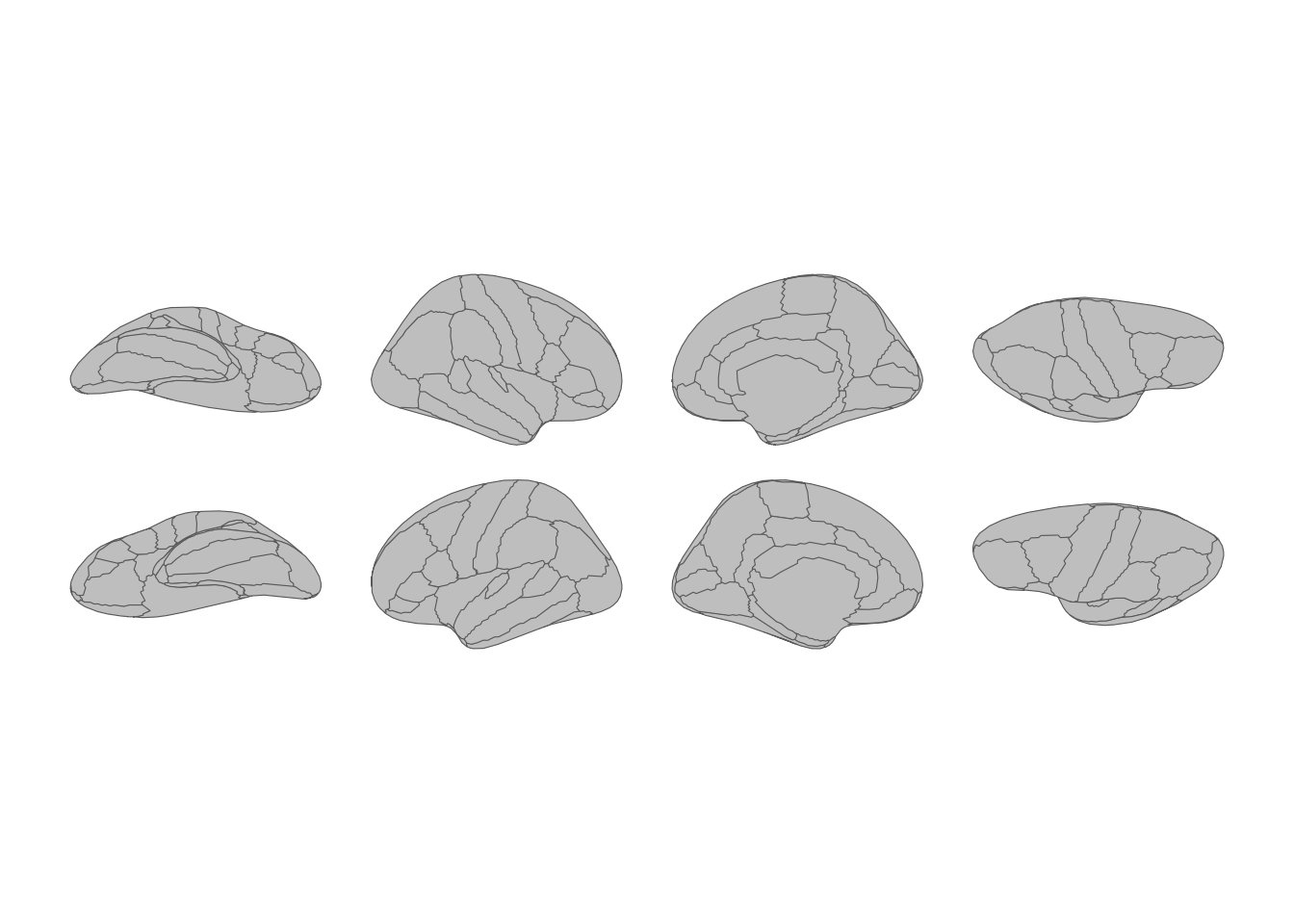
Cortical atlas with hemispheres as rows and views as columns.
Flip the formula to transpose the layout:
ggplot() +
geom_brain(
atlas = dk(),
position = position_brain(view ~ hemi),
show.legend = FALSE
) +
theme_void()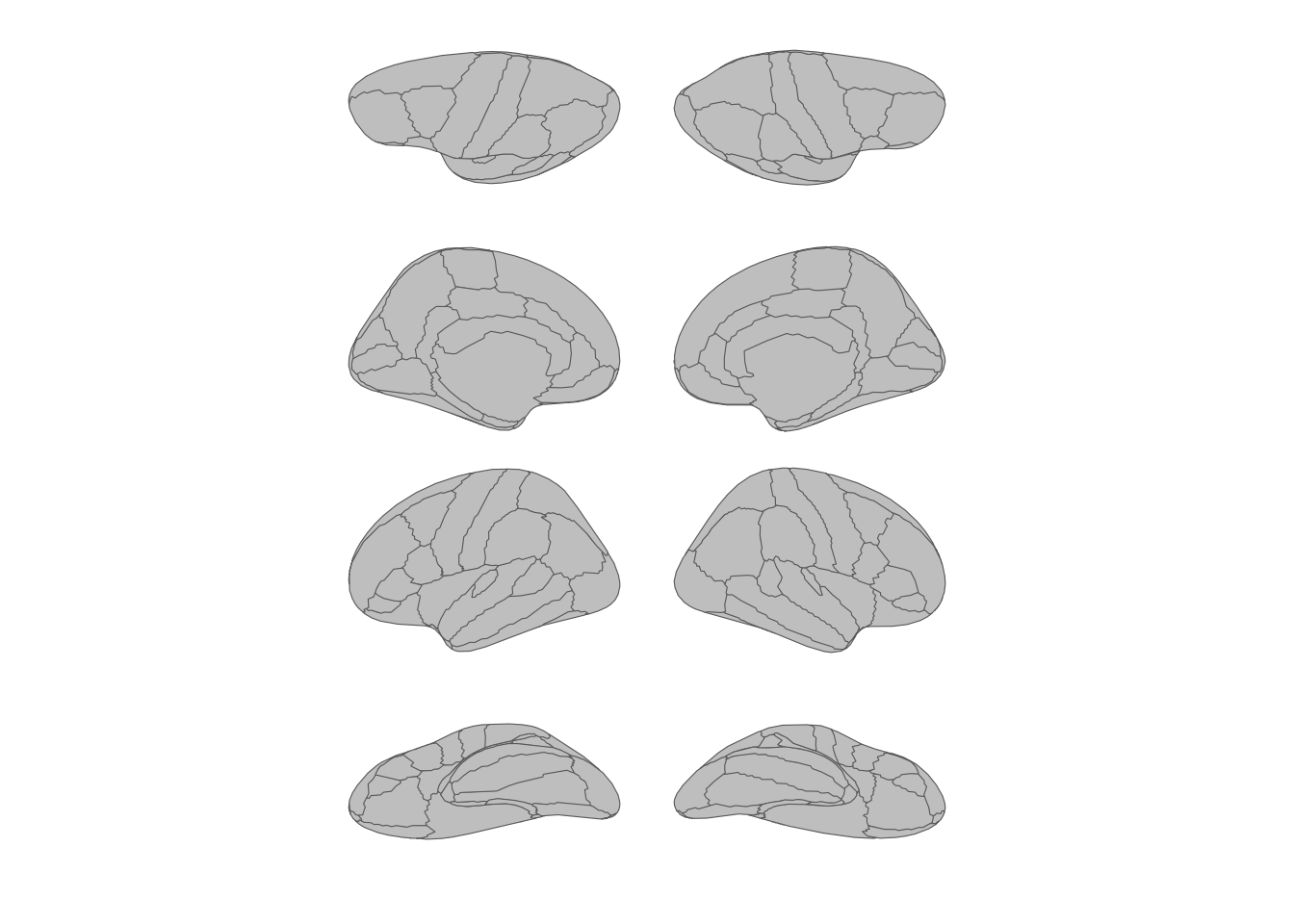
Transposed layout with views as rows and hemispheres as columns.
Stacking all views
Use . with + to collapse everything into a
single row or column. This is handy for compact figures:
ggplot() +
geom_brain(
atlas = dk(),
position = position_brain(. ~ hemi + view),
show.legend = FALSE
) +
theme_void()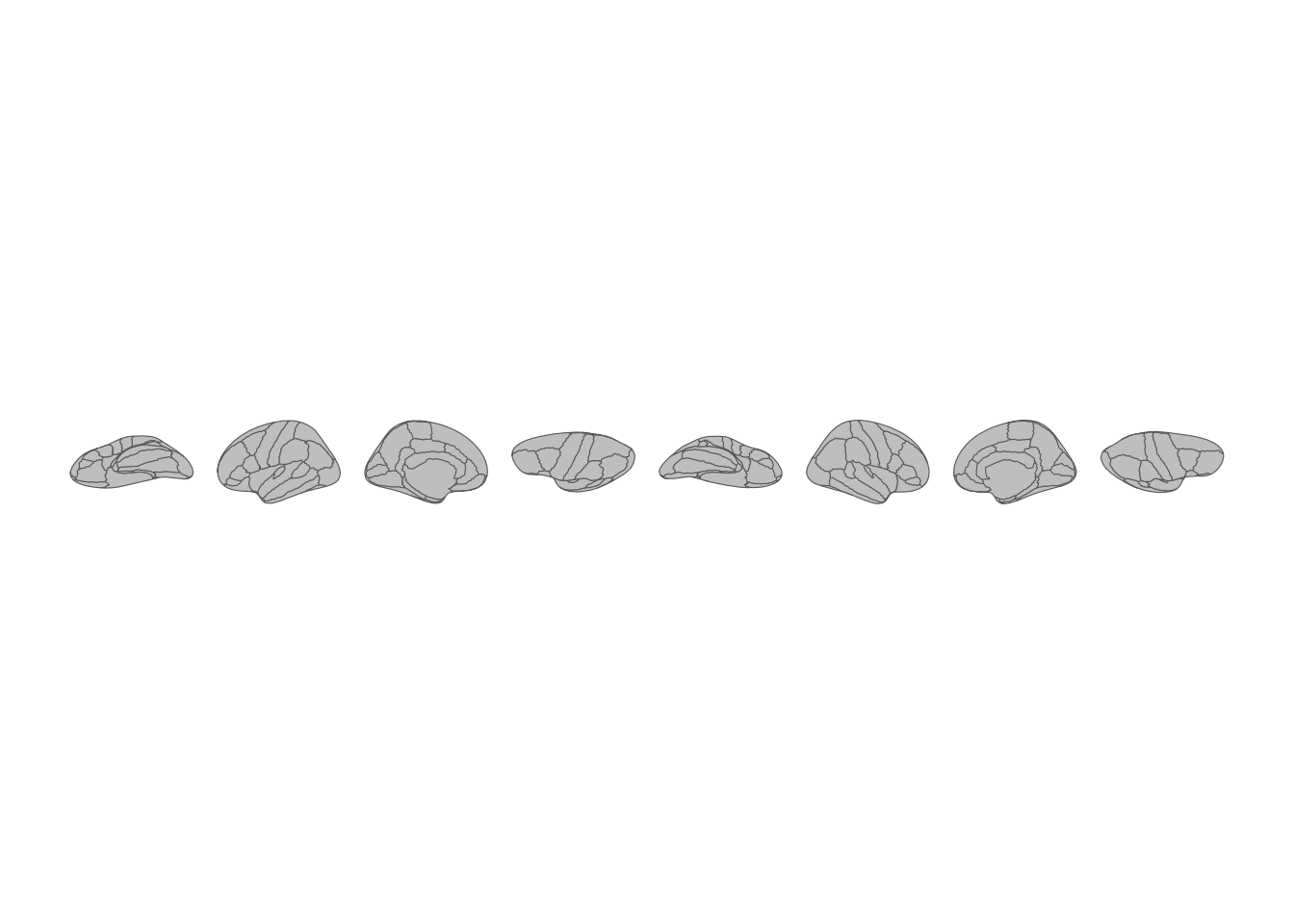
All brain views stacked in a single row.
ggplot() +
geom_brain(
atlas = dk(),
position = position_brain(hemi + view ~ .),
show.legend = FALSE
) +
theme_void()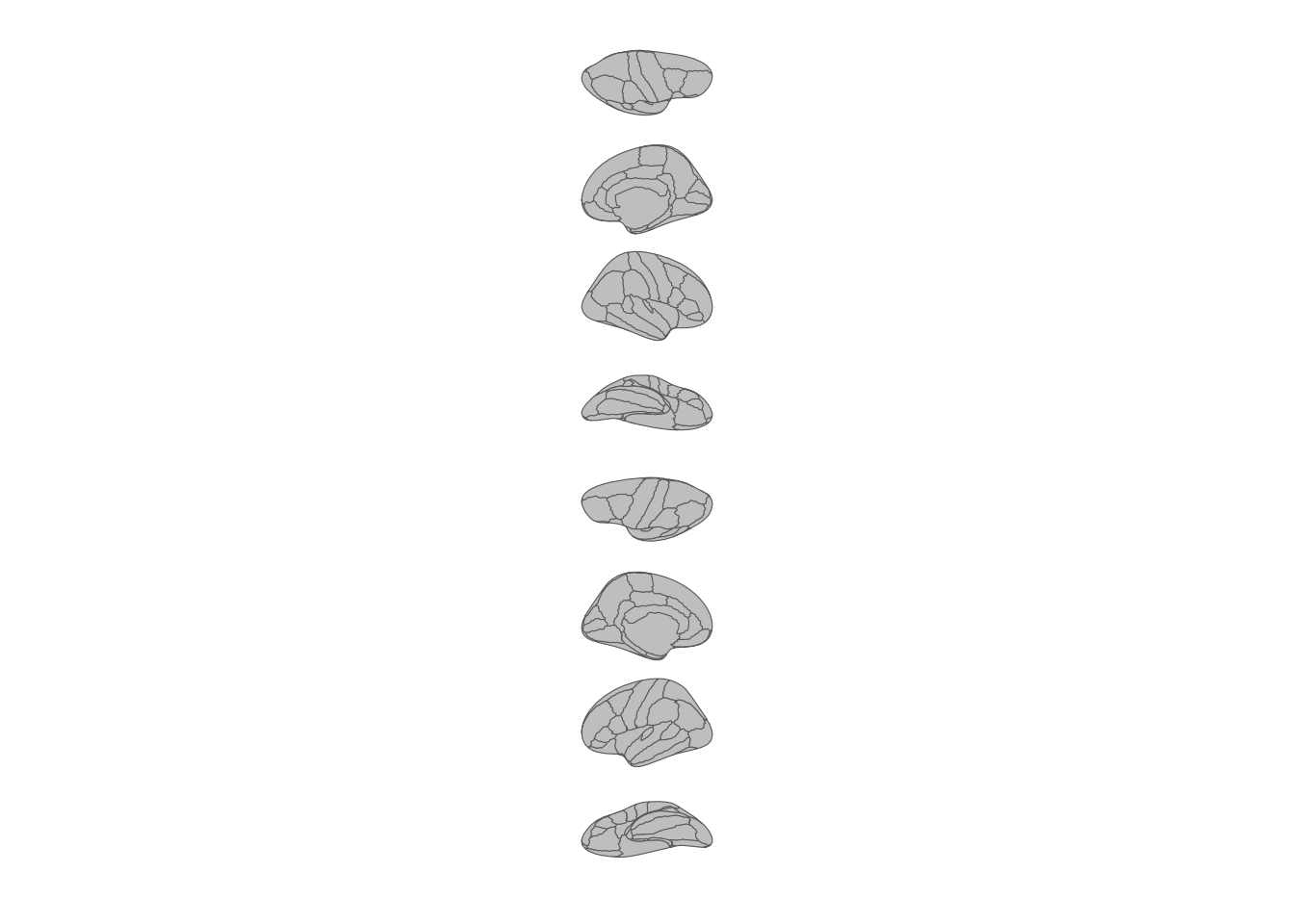
All brain views stacked in a single column.
Subcortical and tract atlases
Subcortical atlases like aseg and tract atlases like
tracula don’t have the hemisphere/view pairing that
cortical atlases do. Their views are individual slices
(e.g. "axial_3", "sagittal"). That opens up a
different set of positioning tools.
Horizontal and vertical
The simplest options. "horizontal" is the default:
ggplot() +
geom_brain(
atlas = aseg(),
position = position_brain("horizontal"),
show.legend = FALSE
) +
theme_void()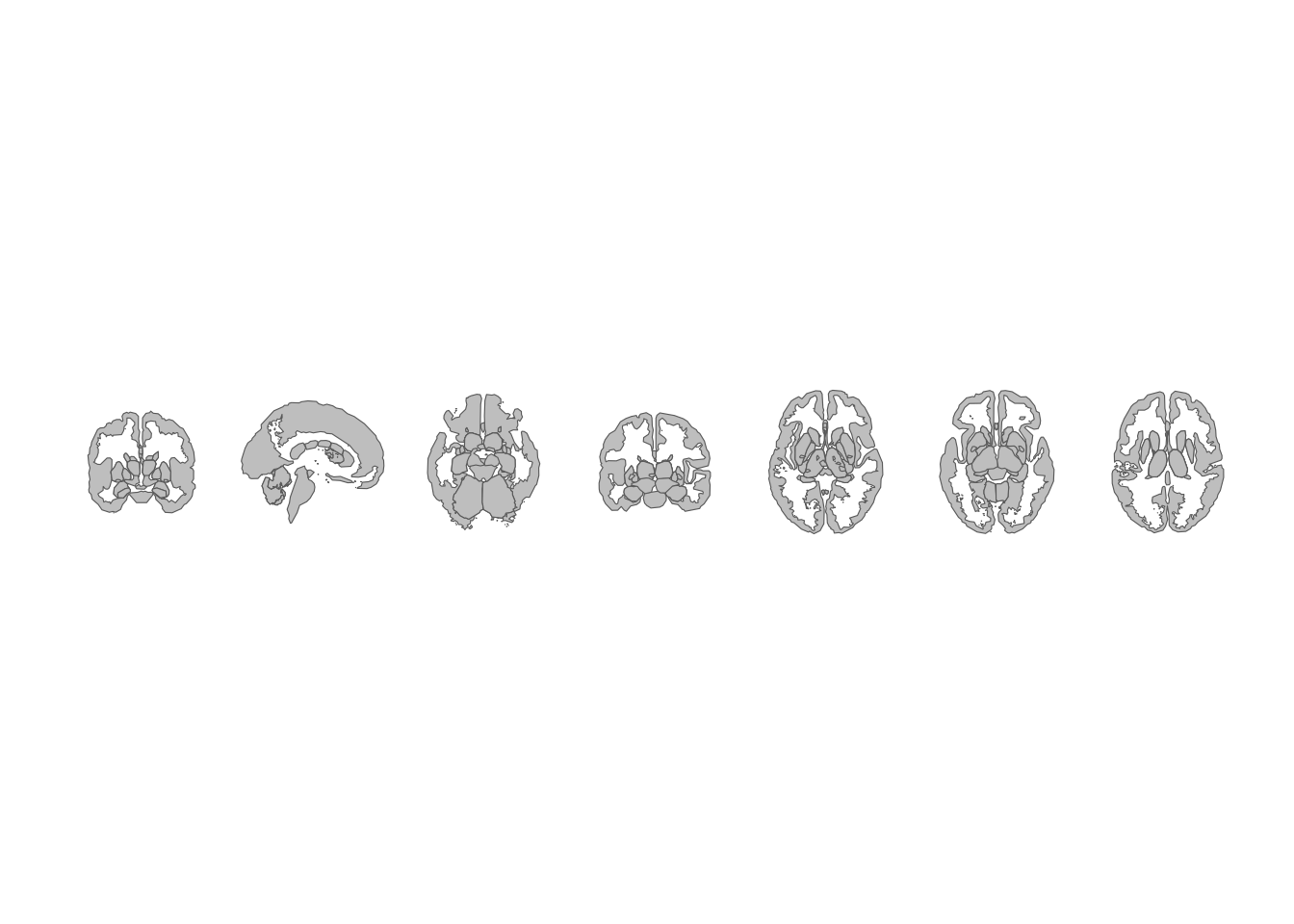
Subcortical atlas views arranged horizontally.
ggplot() +
geom_brain(
atlas = aseg(),
position = position_brain("vertical"),
show.legend = FALSE
) +
theme_void()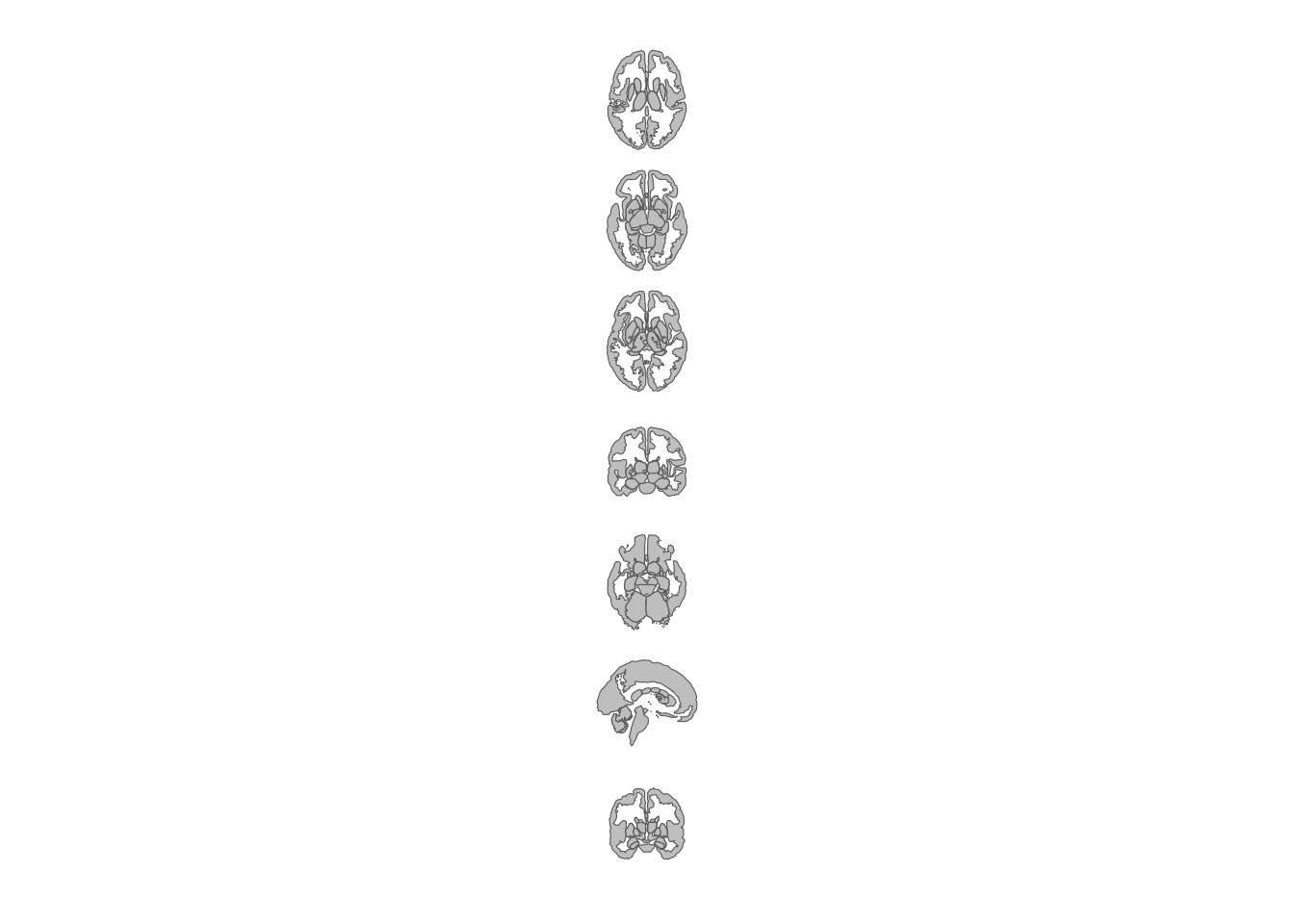
Subcortical atlas views arranged vertically.
Grid layouts
When you have many views, a grid keeps things readable. Specify
nrow, ncol, or both:
ggplot() +
geom_brain(
atlas = aseg(),
position = position_brain(nrow = 2),
show.legend = FALSE
) +
theme_void()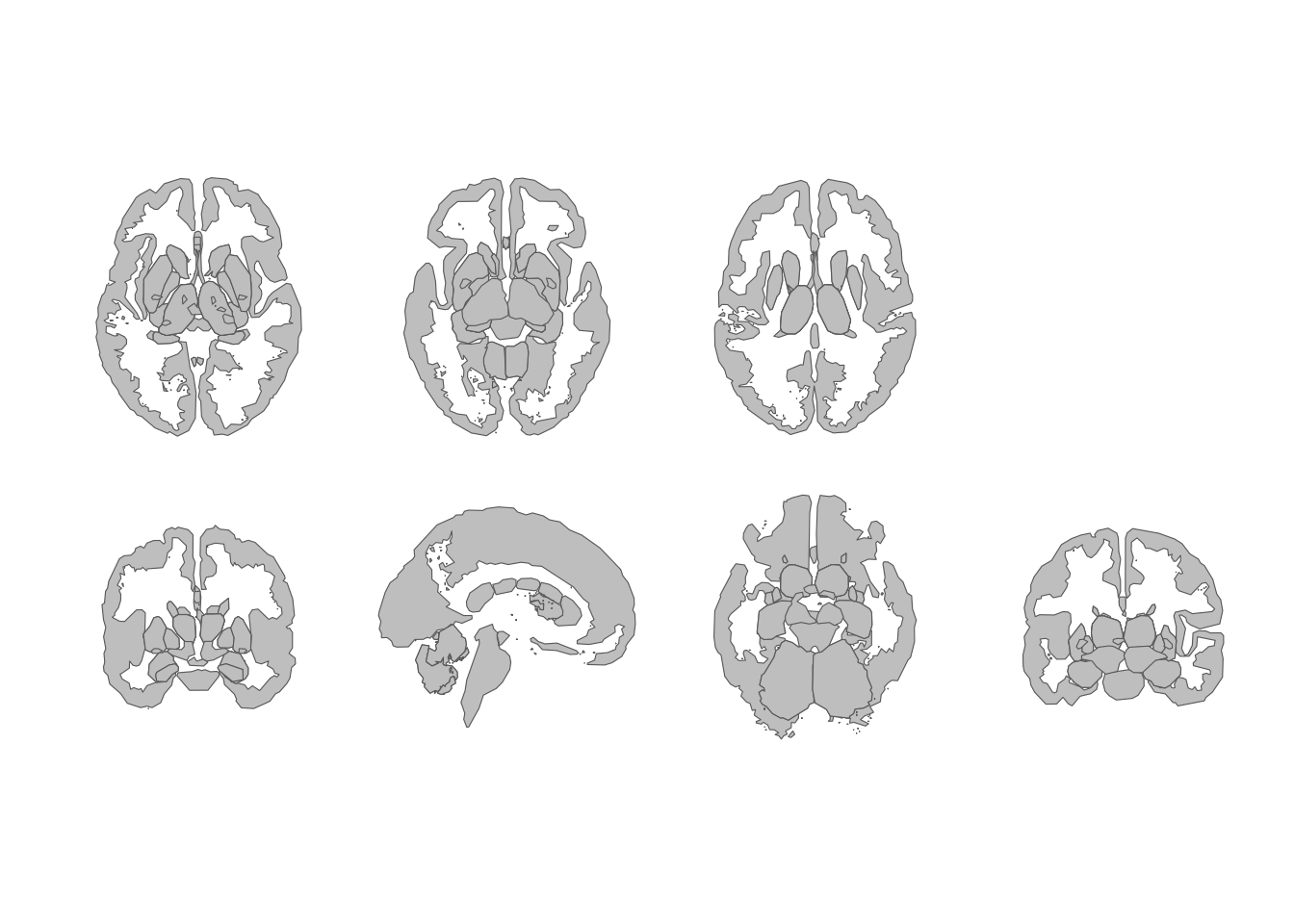
Subcortical atlas views in a two-row grid.
ggplot() +
geom_brain(
atlas = aseg(),
position = position_brain(ncol = 3),
show.legend = FALSE
) +
theme_void()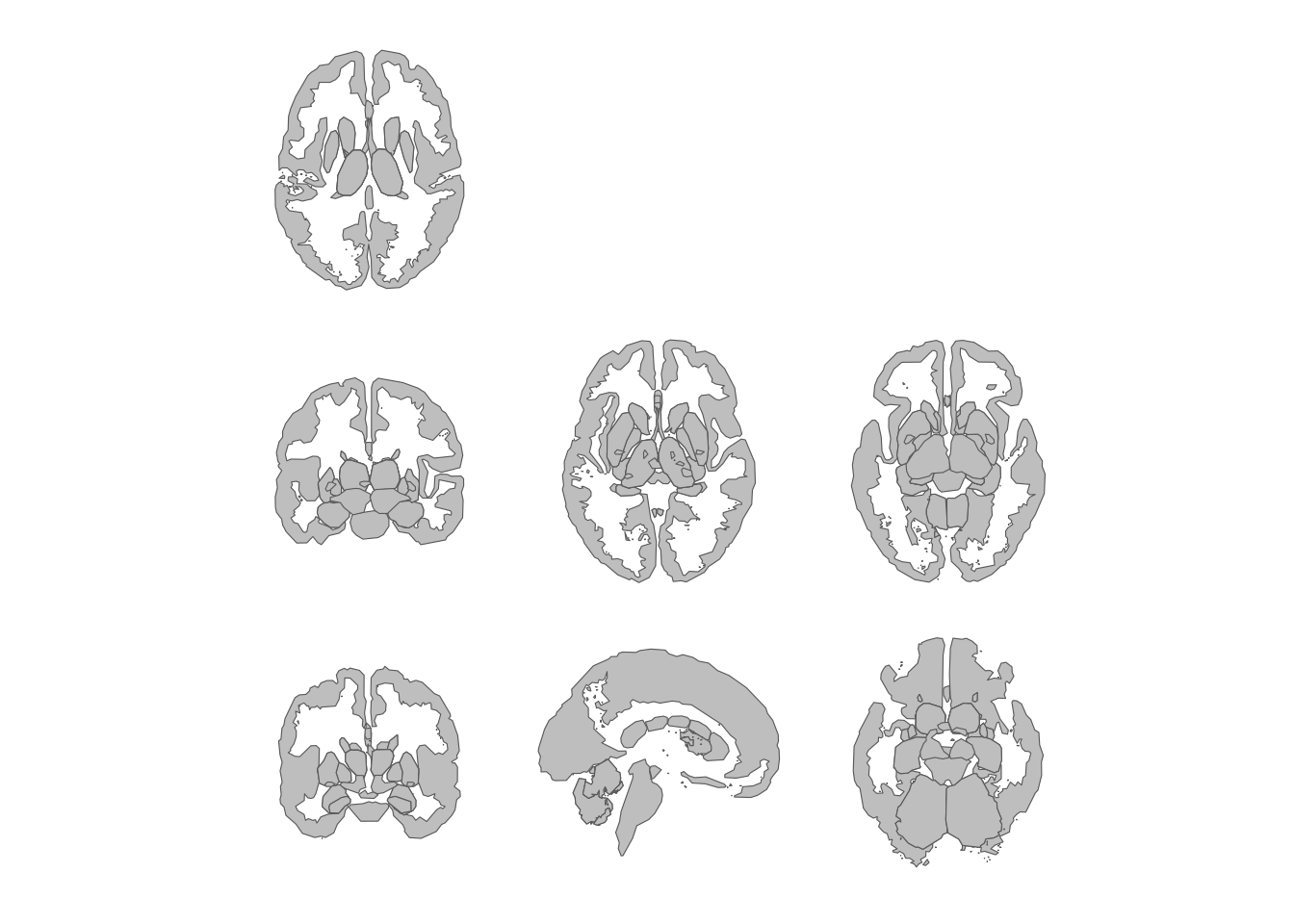
Subcortical atlas views in a three-column grid.
Picking specific views
The views parameter lets you select which views to
include and in what order. Check what’s available with
ggseg.formats::atlas_views():
ggseg.formats::atlas_views(aseg())
#> [1] "axial_3" "axial_4" "axial_5" "axial_6" "coronal_1" "coronal_2"
#> [7] "sagittal"
ggplot() +
geom_brain(
atlas = aseg(),
position = position_brain(
views = c("sagittal", "axial_3", "coronal_3")
),
show.legend = FALSE
) +
theme_void()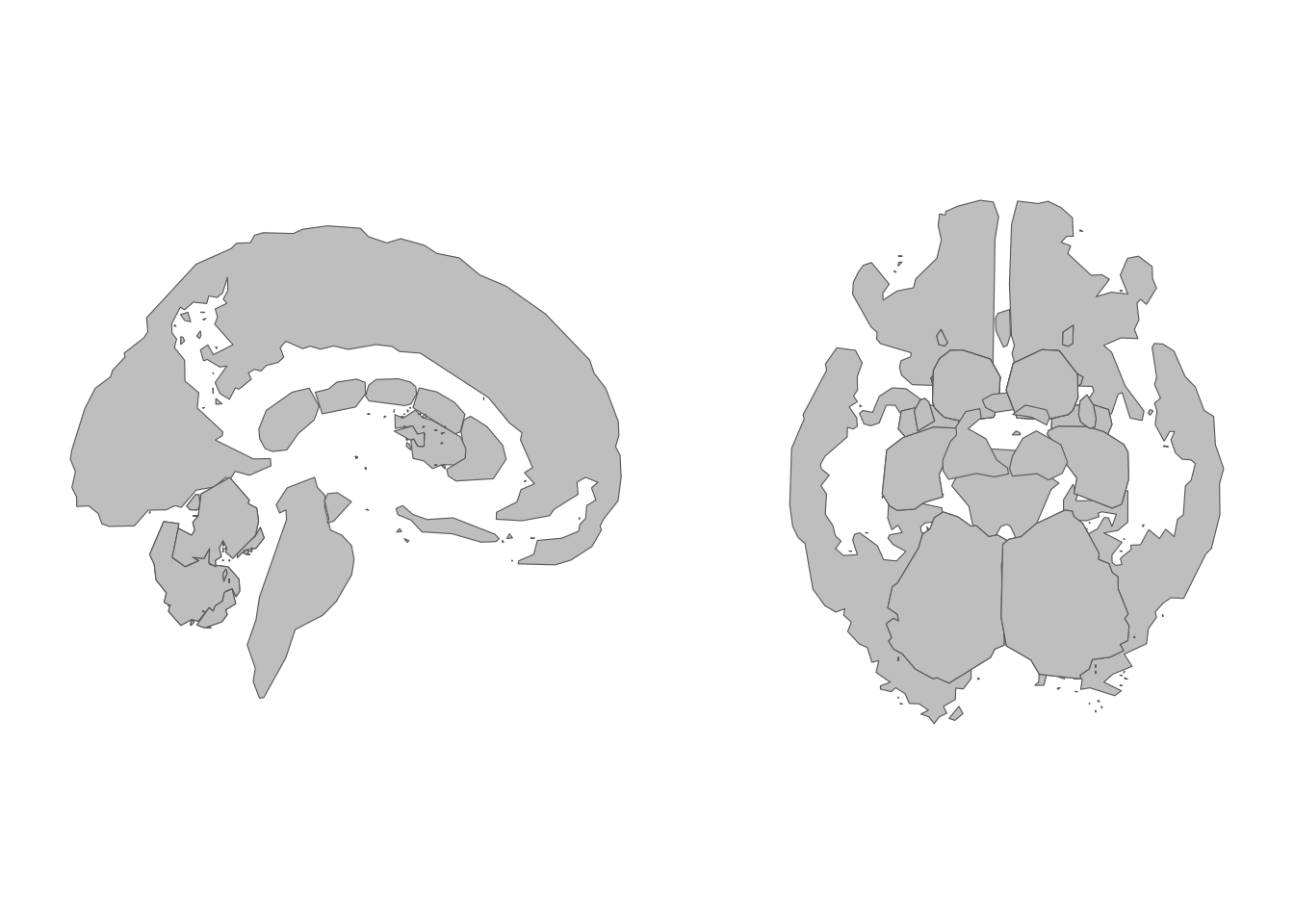
A subset of subcortical views selected by name.
Combine views with nrow or
ncol for a custom grid:
ggplot() +
geom_brain(
atlas = aseg(),
position = position_brain(
views = c("sagittal", "axial_3", "axial_5", "coronal_3"),
nrow = 2
),
show.legend = FALSE
) +
theme_void()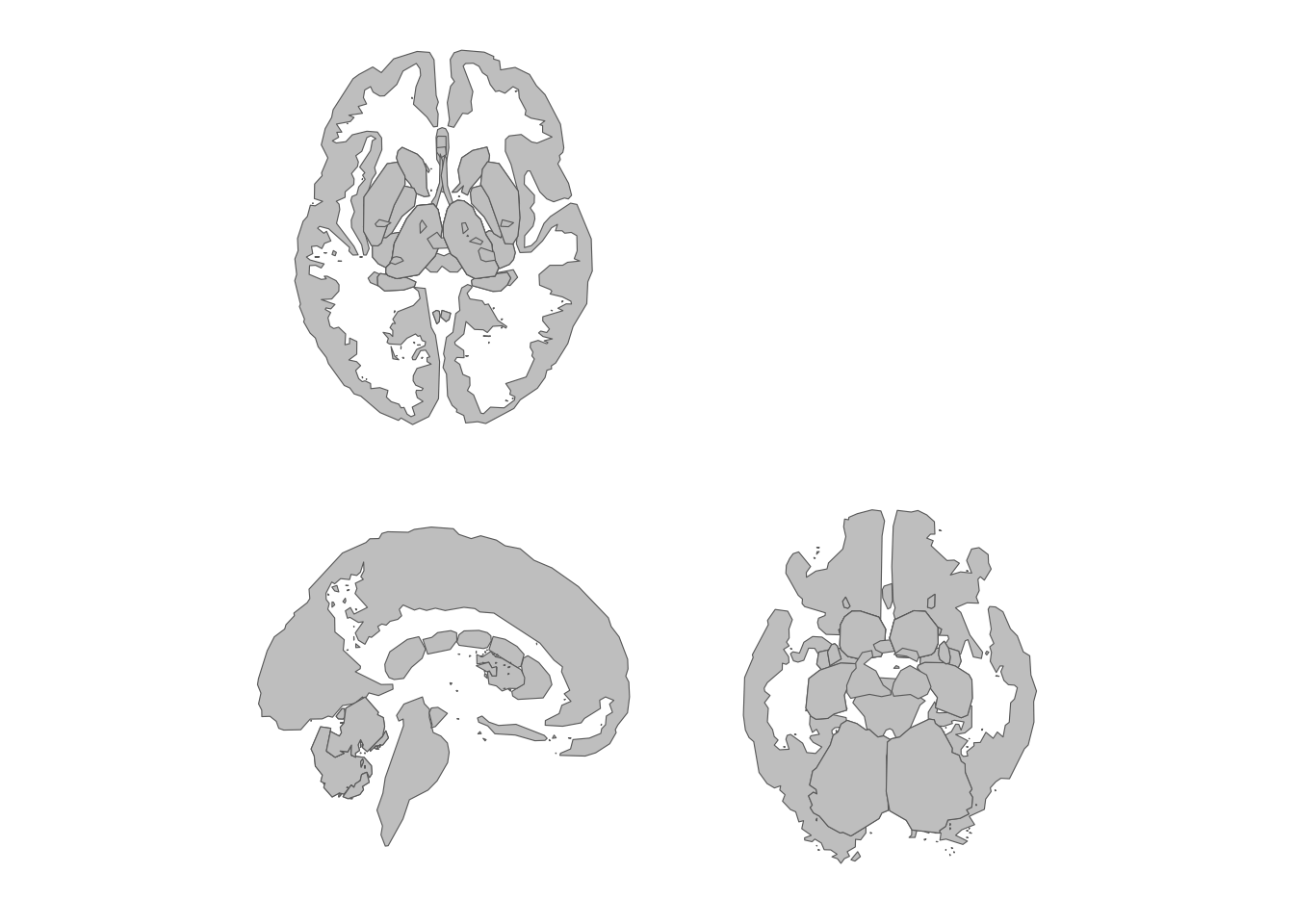
Custom two-row grid with selected subcortical views.
Grouping by slice type
The type ~ . formula groups views by their orientation –
all axial slices together, all coronal slices together, and so on. The
type is extracted from the view name (everything before the first
underscore):
ggplot() +
geom_brain(
atlas = aseg(),
position = position_brain(type ~ .),
show.legend = FALSE
) +
theme_void()Pre-processing with reposition_brain()
If you’re using the geom_sf() workflow instead of
geom_brain(), use reposition_brain() to
rearrange the data before plotting. It takes the same arguments:
atlas_data <- as.data.frame(aseg())
repositioned <- reposition_brain(
atlas_data,
views = c("sagittal", "axial_3", "coronal_3"),
nrow = 1
)
ggplot(repositioned) +
geom_sf(aes(fill = region), show.legend = FALSE) +
theme_void()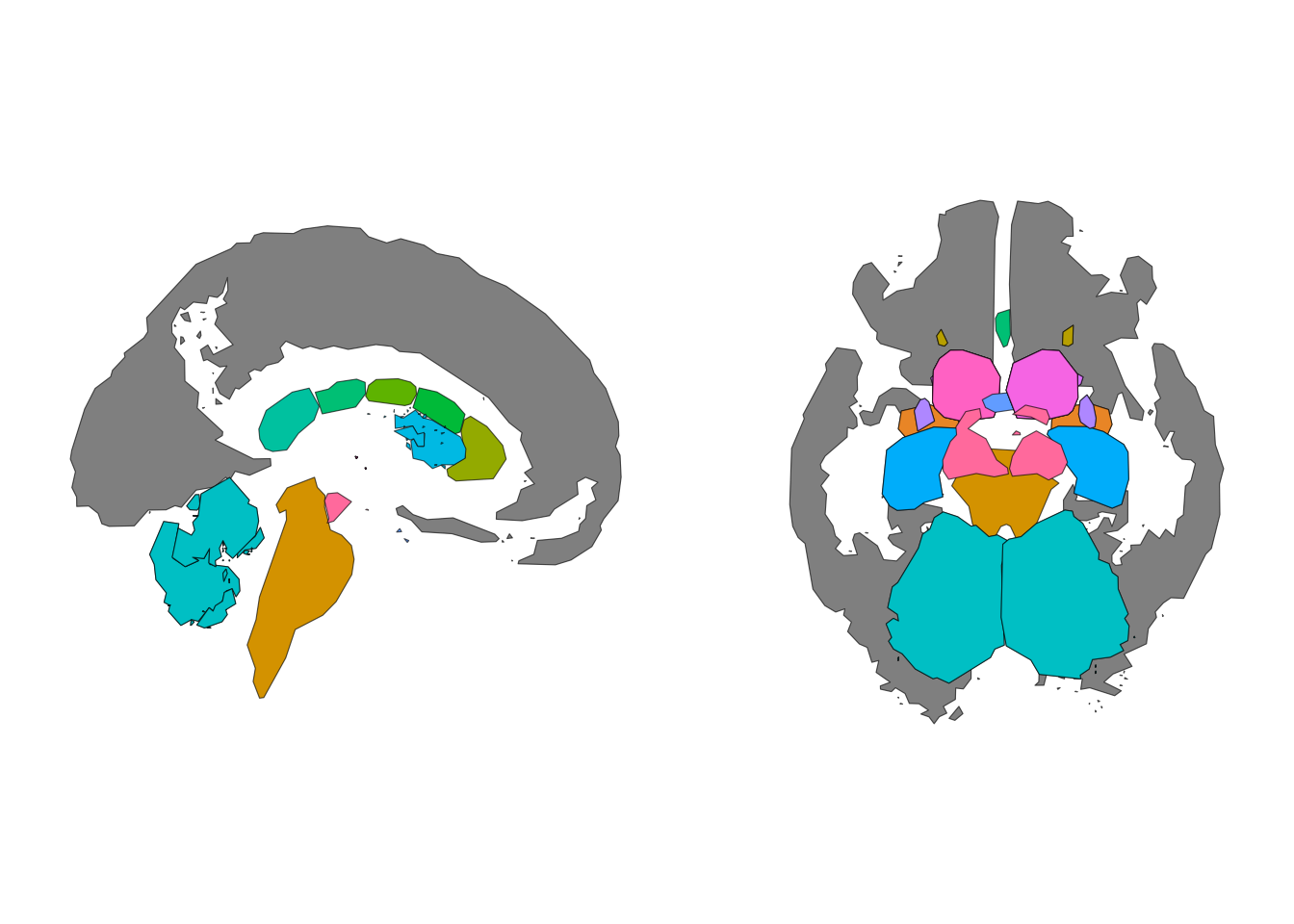
Repositioned subcortical data plotted with geom_sf().
Adding view labels
Use annotate_brain() to label each view with its name.
For cortical atlases the label combines hemisphere and view (e.g. “left
lateral”); for subcortical and tract atlases it uses the view name
directly.
Store the position_brain() specification in an object so
both layers share the same layout:
pos <- position_brain(hemi ~ view)
ggplot() +
geom_brain(atlas = dk(), position = pos, show.legend = FALSE) +
annotate_brain(atlas = dk(), position = pos) +
theme_void()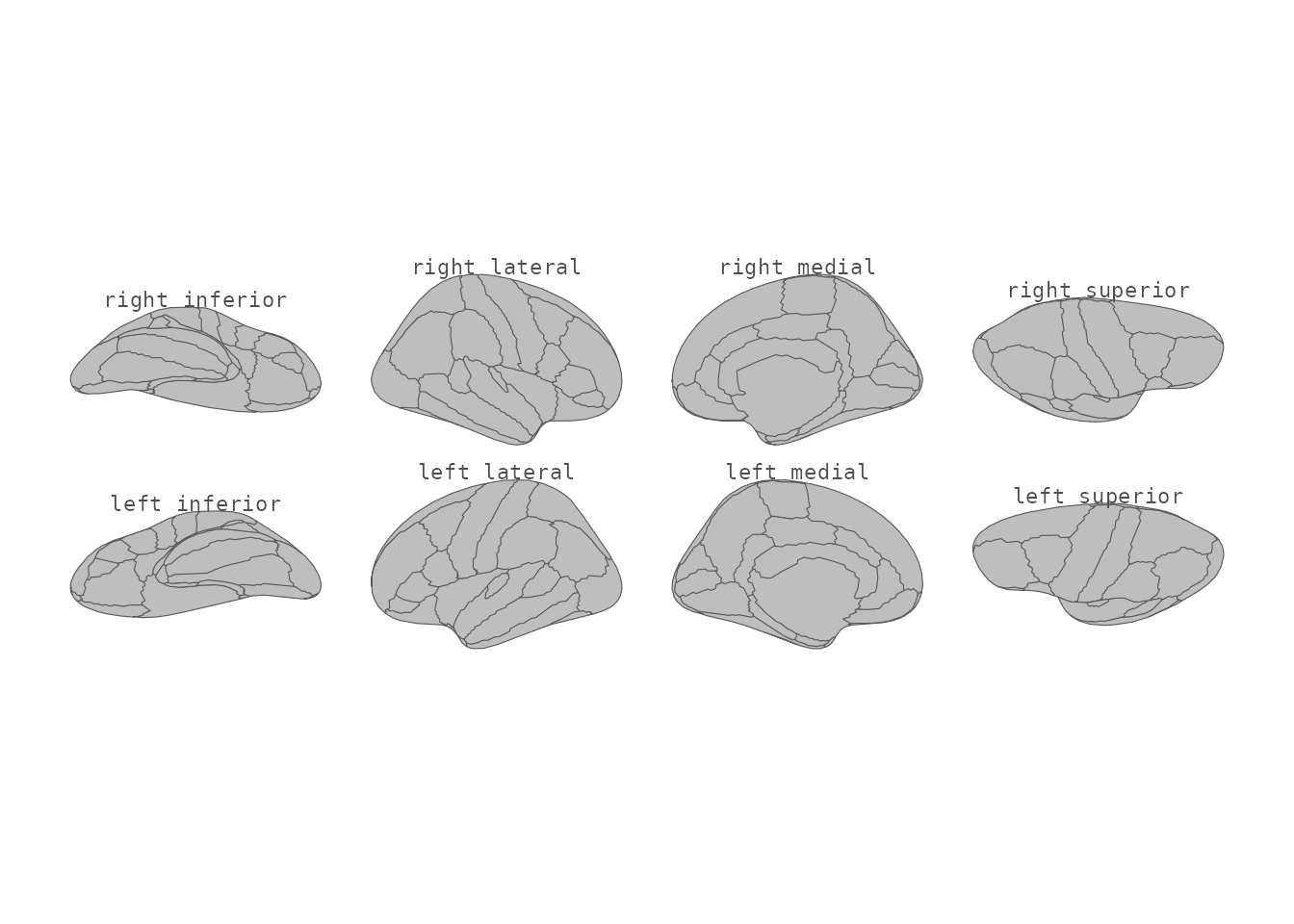
Cortical atlas with view labels.
It works with any positioning — horizontal, vertical, grid, and formula layouts:
pos <- position_brain(nrow = 2)
ggplot() +
geom_brain(atlas = aseg(), position = pos, show.legend = FALSE) +
annotate_brain(atlas = aseg(), position = pos) +
theme_void()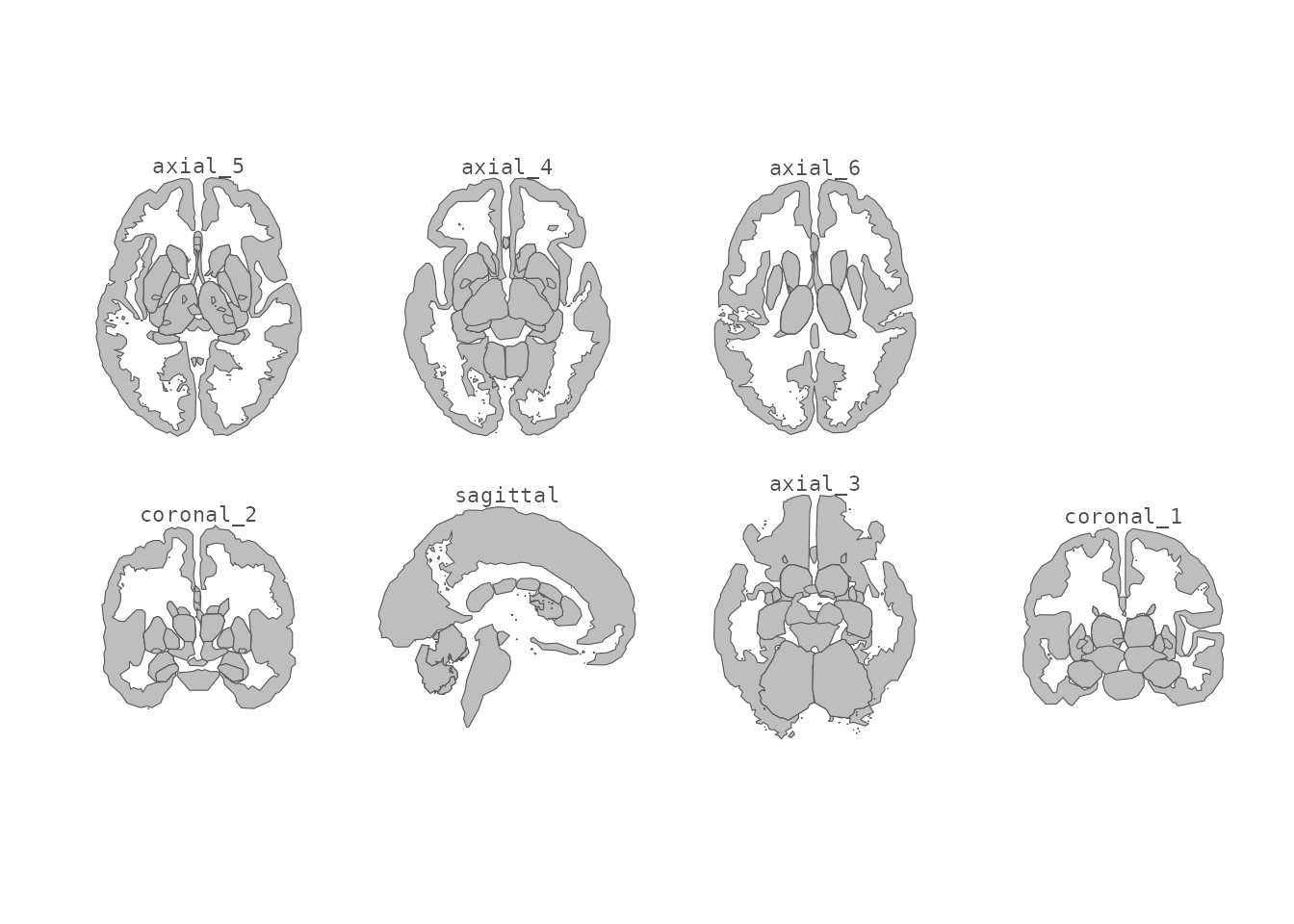
Subcortical atlas with view labels in a two-row grid.
Text appearance is customisable through standard
annotate() arguments:
ggplot() +
geom_brain(atlas = dk(), show.legend = FALSE) +
annotate_brain(
atlas = dk(),
size = 2.5,
colour = "grey50",
fontface = "italic"
) +
theme_void()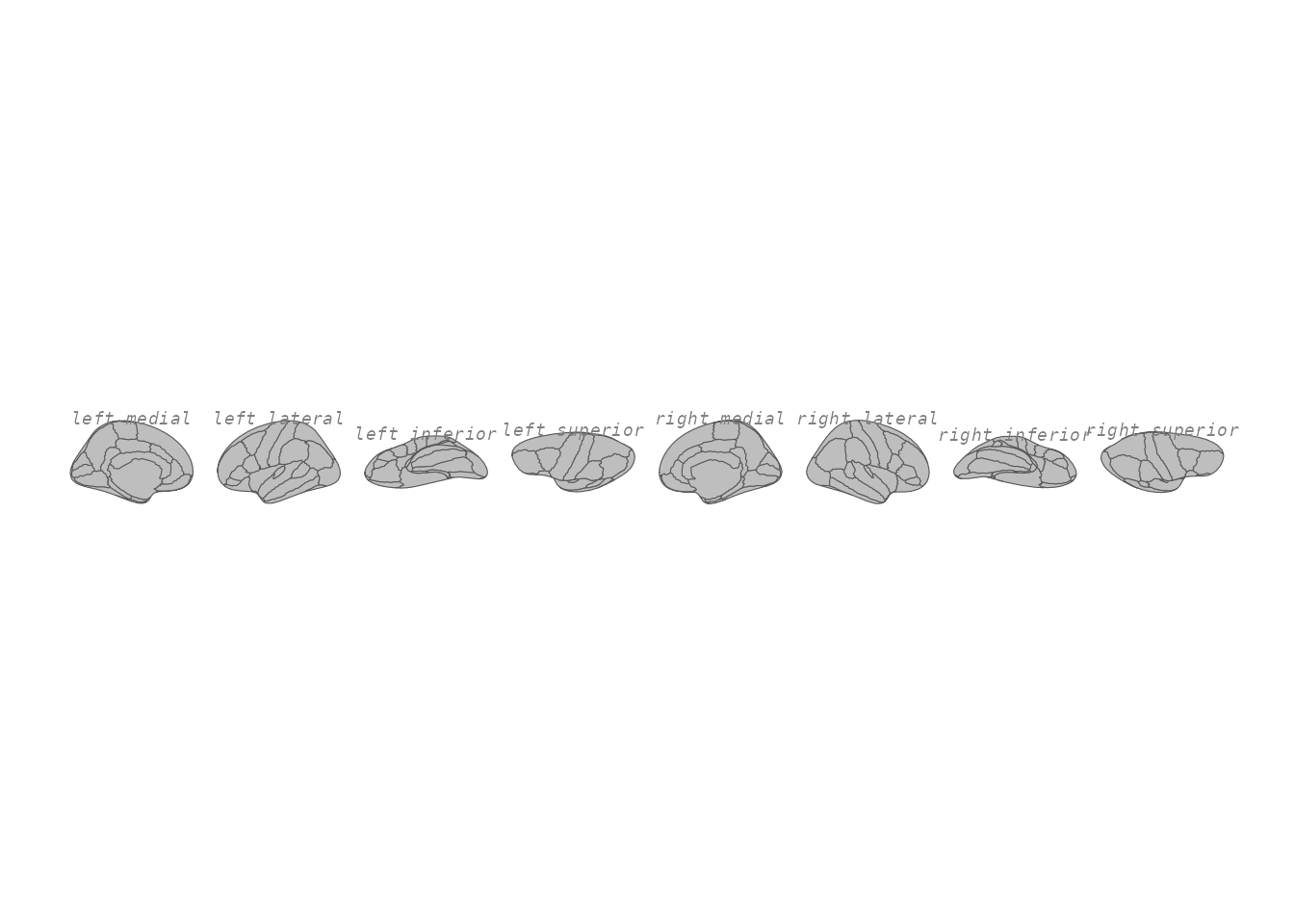
View labels with custom styling.
