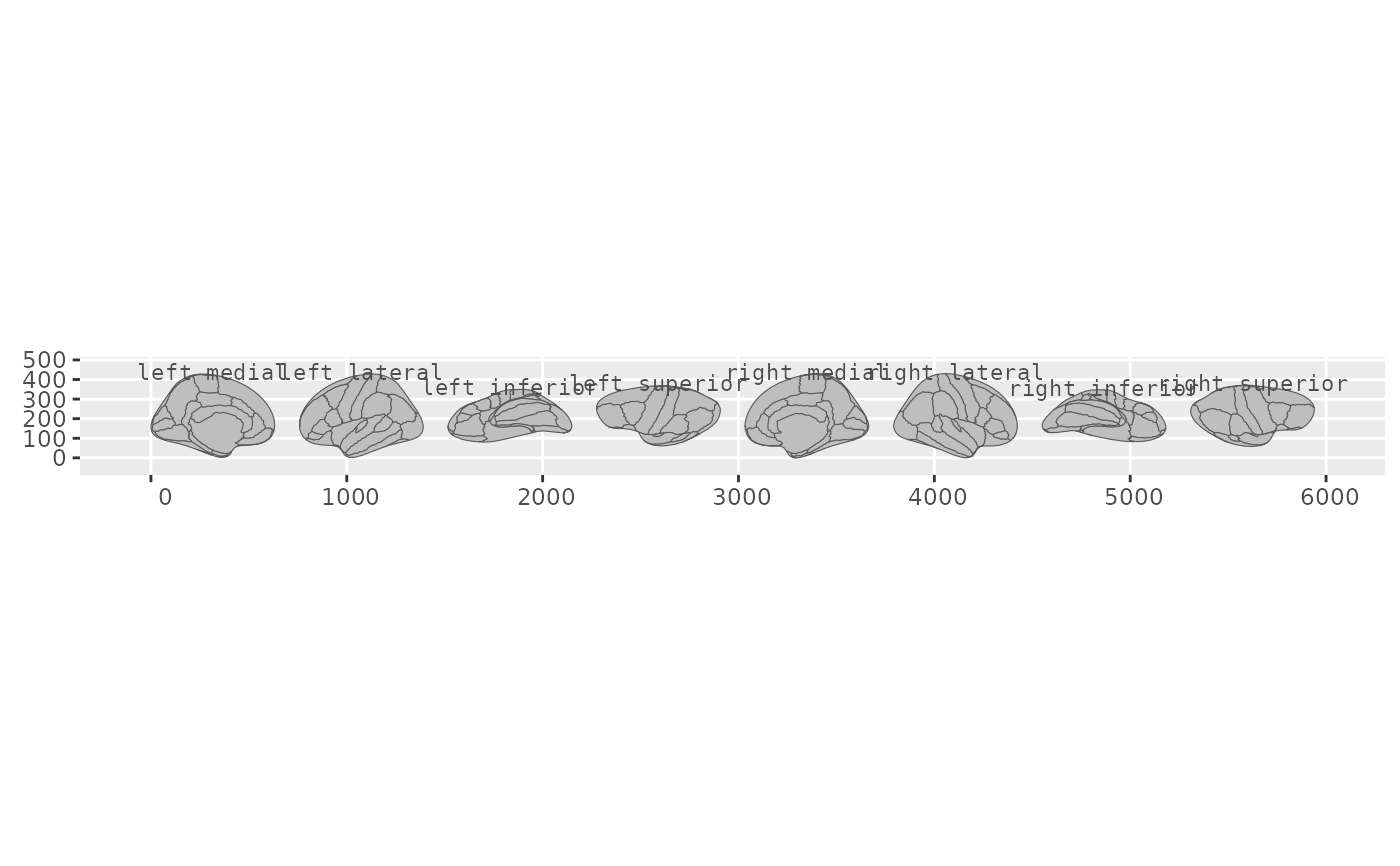
Annotates each brain view with a text label positioned above the view's bounding box. For cortical atlases, labels show hemisphere and view (e.g., "left lateral"). For subcortical and tract atlases, labels show the view name directly (e.g., "axial_1", "sagittal").
Usage
annotate_brain(
atlas,
position = position_brain(),
hemi = NULL,
view = NULL,
size = 3,
colour = "grey30",
family = "mono",
nudge_y = 0,
...
)Arguments
- atlas
A `brain_atlas` object (e.g. `dk()`, `aseg()`).
- position
A [position_brain()] object or position specification matching the one used in [geom_brain()].
- hemi
Character vector of hemispheres to include. If `NULL` (default), all hemispheres are included.
- view
Character vector of views to include. If `NULL` (default), all views are included.
- size
Text size in mm (default: `3`).
- colour
Text colour (default: `"grey30"`).
- family
Font family (default: `"mono"`).
- nudge_y
Additional vertical offset for labels (default: `0`).
- ...
Additional arguments passed to [ggplot2::annotate()].
Details
Labels respect the repositioning done by [position_brain()], so the same `position` argument should be passed to both [geom_brain()] and `annotate_brain()`.
Examples
library(ggplot2)
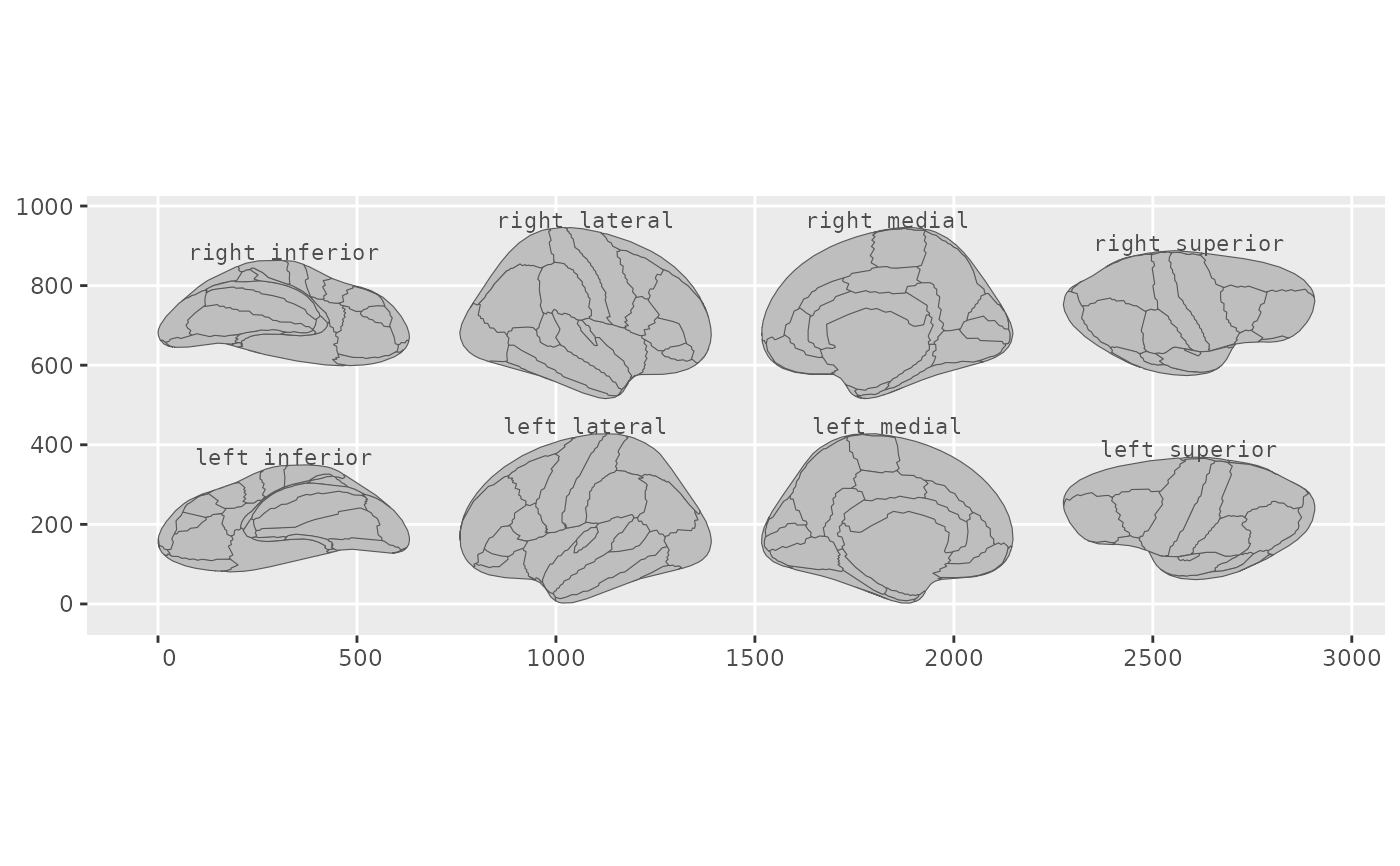
pos <- position_brain(hemi ~ view)
ggplot() +
geom_brain(atlas = dk(), position = pos, show.legend = FALSE) +
annotate_brain(atlas = dk(), position = pos)
 ggplot() +
geom_brain(atlas = dk(), show.legend = FALSE) +
annotate_brain(atlas = dk())
ggplot() +
geom_brain(atlas = dk(), show.legend = FALSE) +
annotate_brain(atlas = dk())